James Foadi(a) and Pierre Aller(b)
(a) Imperial College London (j.foadi@imperial.ac.uk) , Diamond Light Source Ltd (james.foadi@diamond.ac.uk)
(b) Diamond Light Source Ltd (pierre.aller@diamond.ac.uk
Main aim of this tutorials is to get users acquainted with the program BLEND (Foadi et al, 2013) and to show how it can be used effectively to obtain complete and redundant data sets out of several partial or complete ones. The reader interested in exploring pros and cons of using data from multiple crystals can refer, for instance, to Liu et al. (2011), Giordano et al. (2012), Axford et al. (2012), Hanson et al. (2012).
WARNING!!! All merging statistics included in this tutorial have been calculated using POINTLESS and AIMLESS. These CCP4 programs are periodically subject to small changes. It is therefore possible that some of the merging statistics contained in this tutorial appear slightly different from what the user obtains with his/her version of CCP4. These differences are very minimal and should be considered as primarily due to different numerical or algorithmical approximations.
For convenience all tutorials can be carried out inside a same directory and specific results can be collected in separate sub-directories. We will assume that "blend_tutorial" is the name of the overall directory and that it has been created in the $HOME area ("/home/james" in our case). Different directory names and locations can be used, but appropriate modifications will have to be applied to the paths displayed in this document. Data downloaded using the above links, when unpacked, yield a directory named "data". For the rest of the tutorials this directory can be moved inside "blend_tutorial". There are three groups of data in the sub-directory “data”. One group is formed by crystals of insulin, the second by crystals of lysozyme and the third by lysozyme soaked with bromine.
The first tutorial will be illustrated with and without use of the CCP4 interface. Other tutorials will be illustrated either with or without interface, but the user should have no problem re-running them as it suits her / him best.
Easy tutorial to get you started with BLEND
a) Using CCP4I
The CCP4 interface associates a project directory to an already-existing physical directory in your computer file system. Thus we need to create a directory before anything else. Let's call this directory “insulin” (it is assumed you are in the blend_tutorials directory):
mkdir insulin
Next we need to associate a project directory called “BLEND_TUTORIAL_INSULIN” to the insulin directory just created. You can do this using “Directories & ProjectDir” in ccp4i. Now that the project directory has been created let us change to the “BLEND_TUTORIAL_INSULIN” one using “Change Project” in ccp4i. We are ready to start practising with BLEND.
Insulin data have been collected at the Diamond Light Source synchrotron and integrated using MOSFLM. Angular rotation range varies from data set to data set, as these have been collected from different crystals. Please, explore some of these integrated data clicking “View Any File” in ccp4i and selecting the “mtz MTZ” field in “File type”.
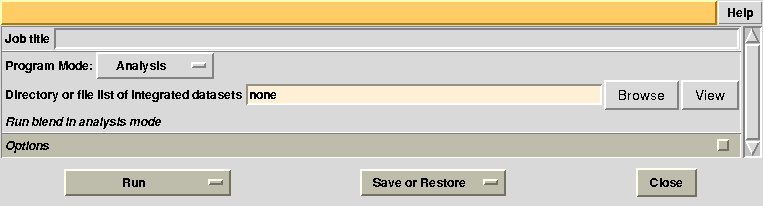
In the CCP4 GUI, BLEND can be found either in the “Data Reduction and Analysis” or the “Program List” section on the left. Once initiated, the BLEND section looks like the following:

To begin with we run BLEND in analysis mode, with the main goal of creating clusters of data that can, potentially, merge well together. After having filled the blank fields with the appropriate paths and keywords, the above interface should resemble this:
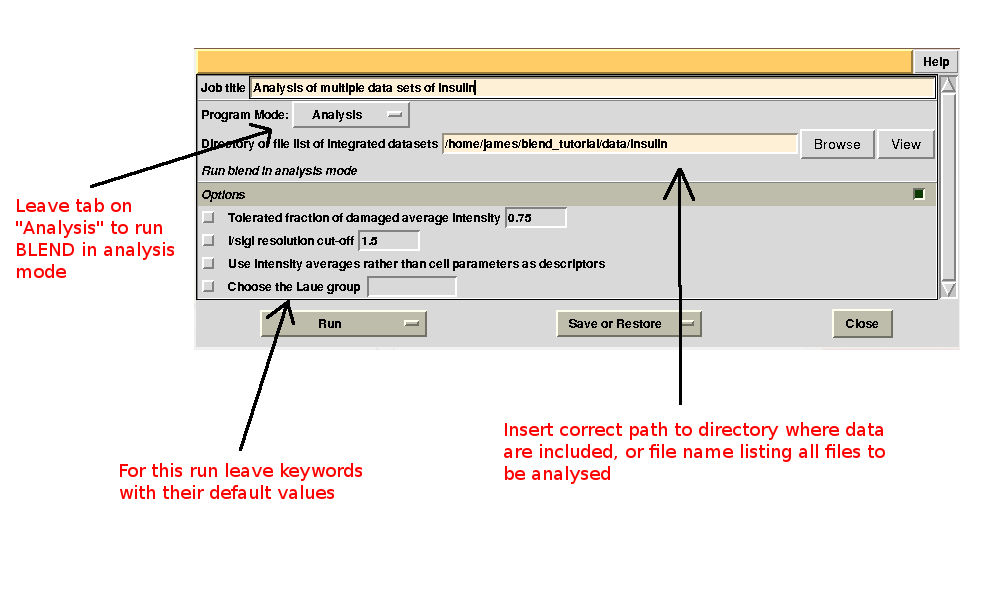
All keywords with their meaning are explained in BLEND documentation. After the run in analysis mode, we can look at the output using ccp4i “View Files from Job → View Job Results (new style)”:
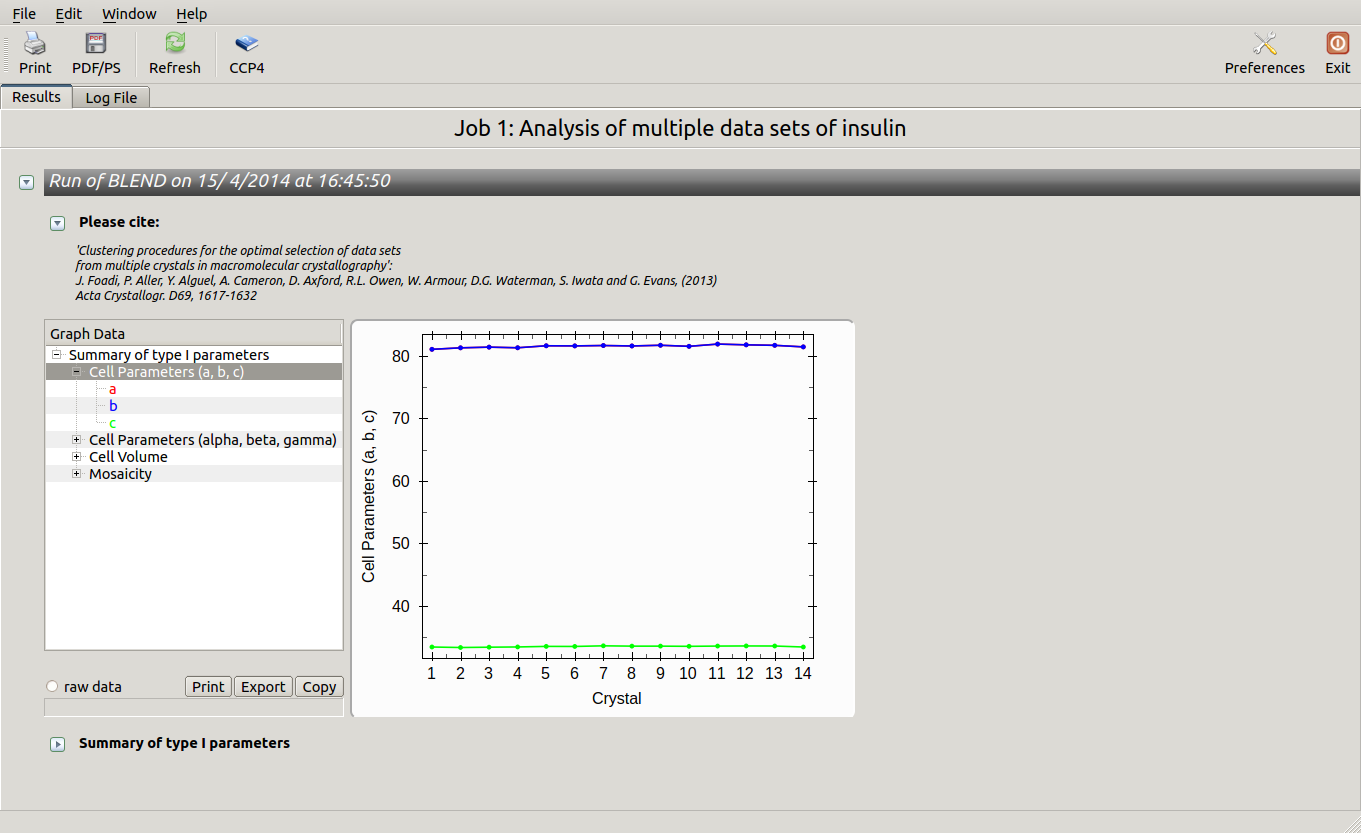
Cell sides, cell angles, cell volume and mosaicity can be viewed for all the 14 data sets included in this insulin tutorial. Numerical values can be explored within the same view. More files have been produced by this run of BLEND in analysis mode. These are listed by selecting “View Files from Job” → “blend_1”:
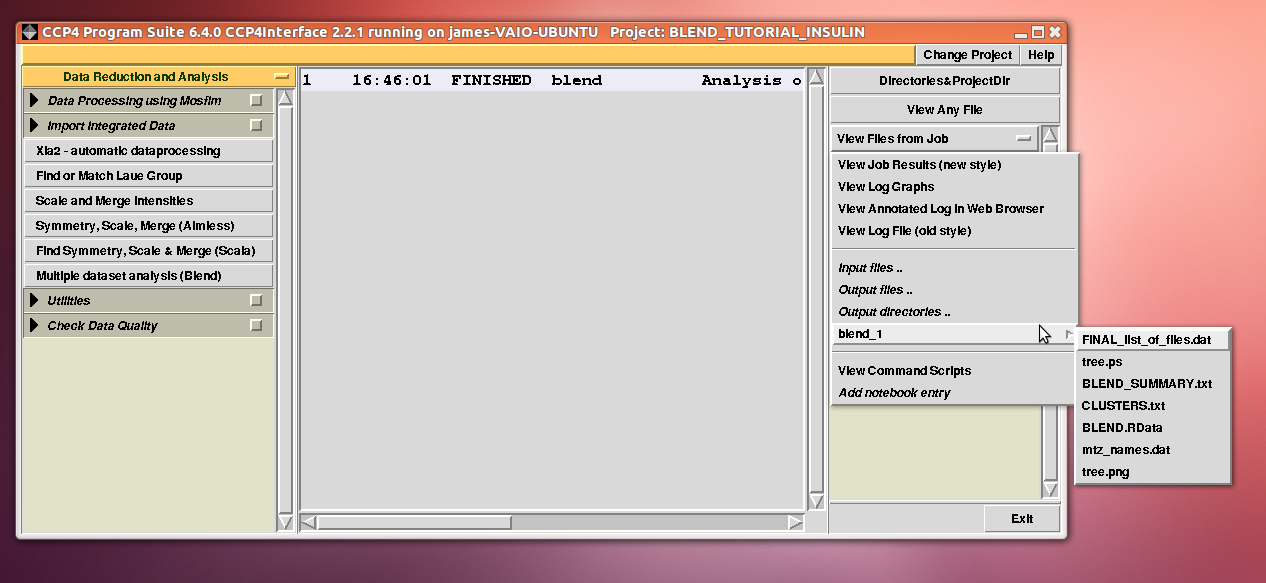
“blend_1” is a directory under the “insulin” directory, containing all files produced by this run of BLEND. The most important among these files is possibly the dendrogram (files “tree.ps” and “tree.png”). Both PNG and PS files display the same dendrogram. They can be viewed using any graphical tool, but, at present, only the PS file can be visualised using the CCP4 GUI:
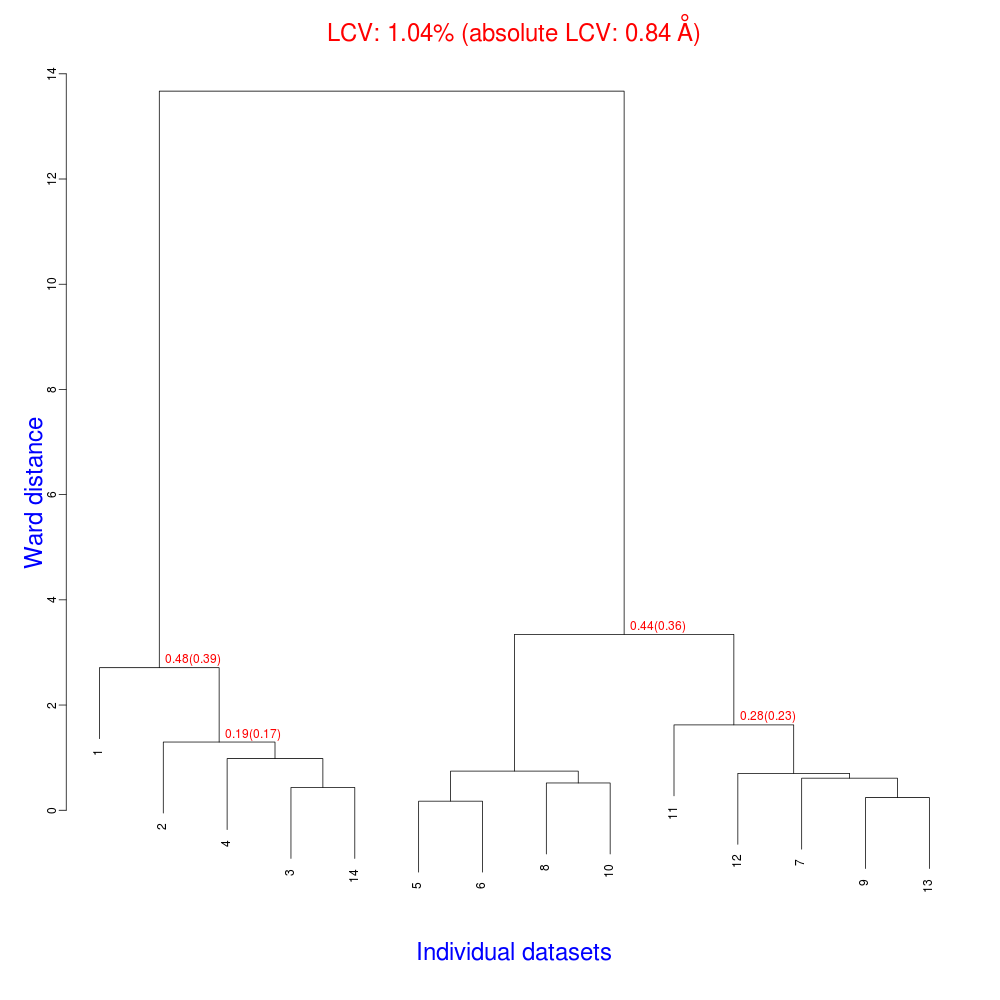
The numbers in red are the Linear Cell Variation (LCV) values (and the associated absolute LCV in Å) for the top five clusters. They give an indication of cell similarity among all crystals included in the specific cluster; thus, ultimately, they can be associated with isomorphism between different data sets. In the above dendrogram we can see two clusters with similar cell variability (0.44% and 0.48%, corresponding to 0.36 Å and 0.39 Å respectively).The variability is more than doubled when these clusters merge into the overall cluster containing all 14 data sets; this is probably an indication of some minor form of non-isomorphism. Tests done so far with BLEND show that some structural dissimilarities can be noticed for values of LCV higher than 1 – 1.5%. It is probably more appropriate to use absolute values in order to make judgments on crystals isomorphism. The reason is that the absolute Linear Cell Variation (aLCV) is expressed in angstroms and, thus, it can immediately give sensible numbers in relation to cell parameters differences.
LCV and aLCV values for all clusters are reported in the file “CLUSTERS.txt”:
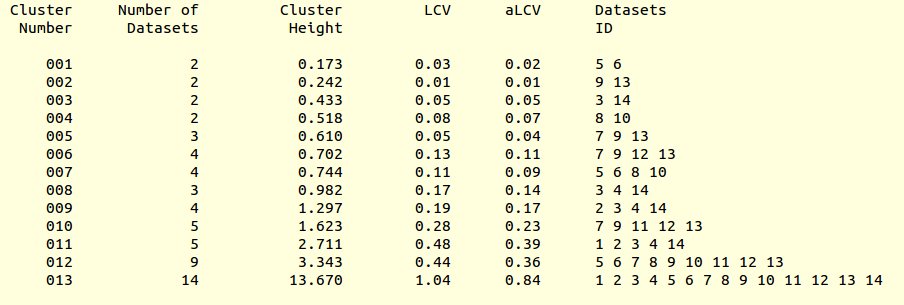
This is, essentially, a numerical version of the dendrogram and, as we will see, is often useful for executions of BLEND in synthesis mode. Another convenient file in directory “blend_1” is “BLEND_SUMMARY.txt”, which is essentially a table reporting cell parameters and other quantities for each data set. The unique serial number assigned by BLEND to each data set can be found in the “FINAL_list_of_files.dat” file. For us this file looks like the following:
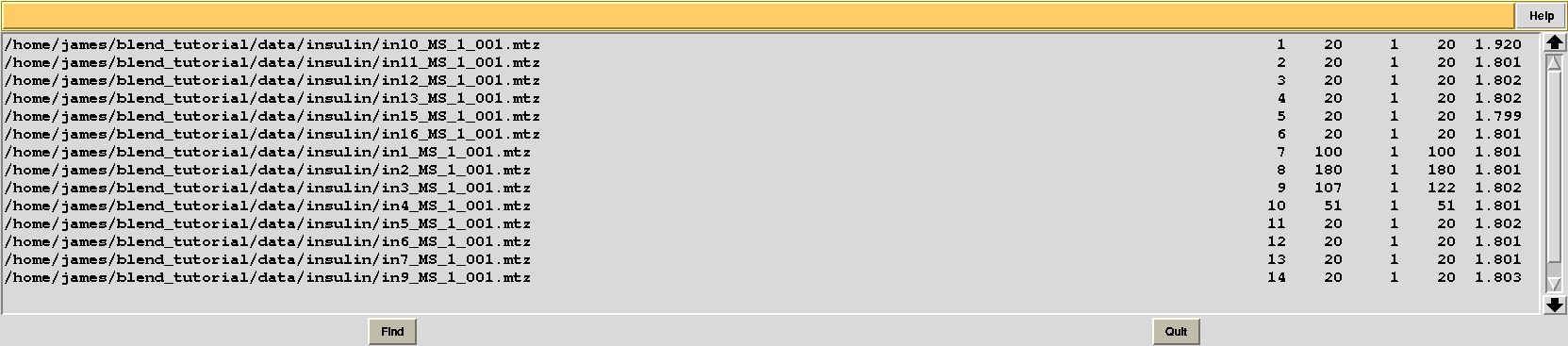
The first column contains full path to all input data files; the second contains the serial number assigned to input files; the fourth and fifth column list initial and final image numbers, while the third lists the last image BLEND will include for all subsequent scaling and merging jobs. Numbers in these column have been calculated through procedures to get rid of radiation damaged images. In the specific case the only data set where images will be discarded is data set 9, for which BLEND suggests to remove from image 108 to image 122. The last column include resolution cuts suggested by BLEND for all subsequent scaling and merging jobs. These are worked out using intensity averages decay with resolution, and are controlled by keyword ISIGI.
All input data files were assigned a serial number; this is not always the case, because sometimes individual data sets are excluded as they contain multiple wedges or other irregular features, and BLEND does not accept them in the group to be further analysed. There might also be reasons for which the user does not want to consider some of the files included in the data directory (for example if they belong to an unwanted experiment). In this case the user could consider file “mtz_names.dat”, also included in directory “blend_1”, to re-run BLEND using less input data. File “mtz_names.dat” looks like the following:
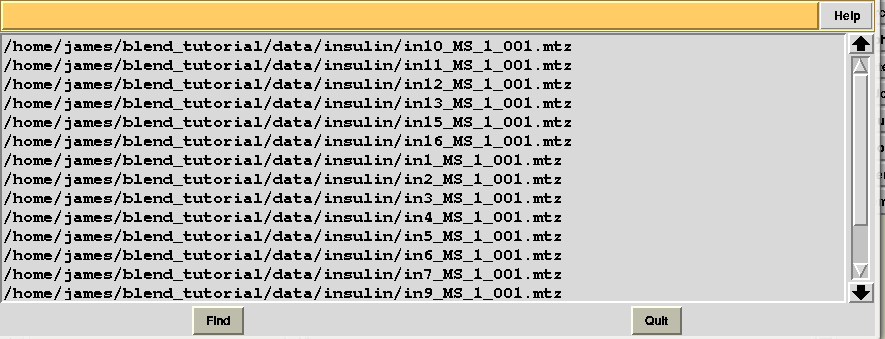
In order to show how to make use of this file suppose we are not any longer interested in the last 4 data sets. We first copy “mtz_names.dat” into a file called “original.dat”, in a newly-created directory called “amended_insulin”:
mkdir amended_insulin
cp insulin/blend_1/mtz_names.dat amended_insulin/original.dat
At this point the last 4 files can be deleted from “original.dat” file which, now, looks like the following:
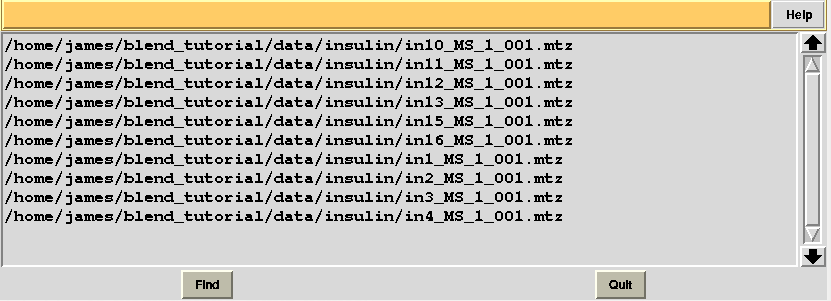
Next, we associate this directory to a new project directory in ccp4i, called “BLEND_TUTORIAL_AMENDED_INSULIN”, and select this project directory. BLEND can now be executed again in analysis mode, this time selecting the “original.dat” file as starting point, rather than the whole directory containing all 14 insulin data sets:
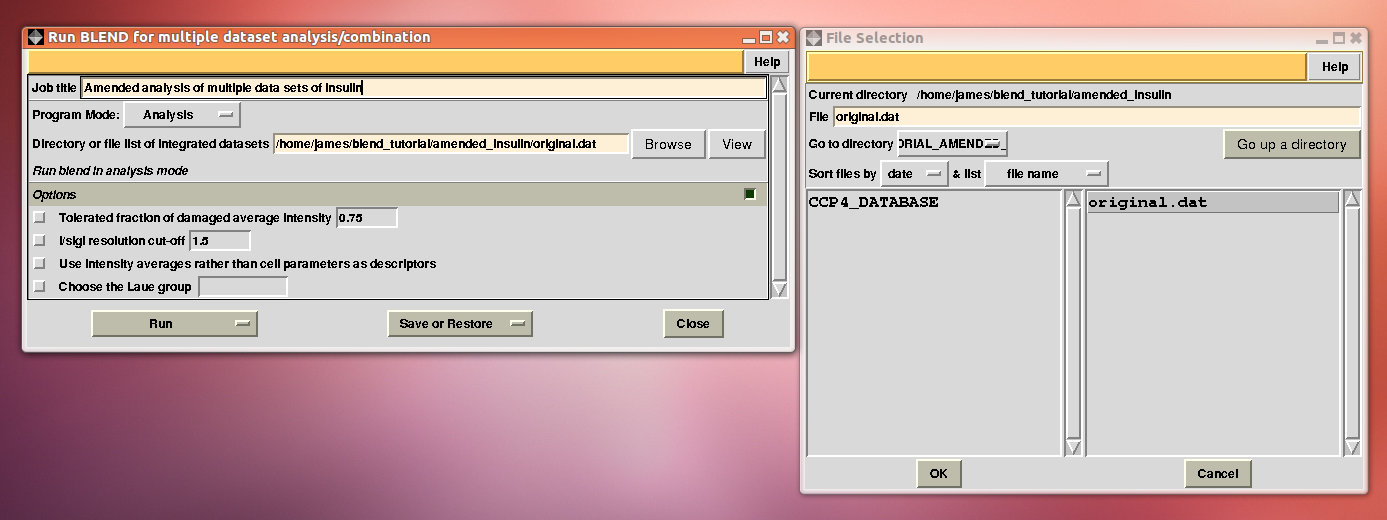
The usual files will be produced (still in directory “blend_1”, but inside directory “amended_insulin”), although this time BLEND will have dealt with 10, rather than 14 data sets, and the dendrogram will look different from the one previously obtained:
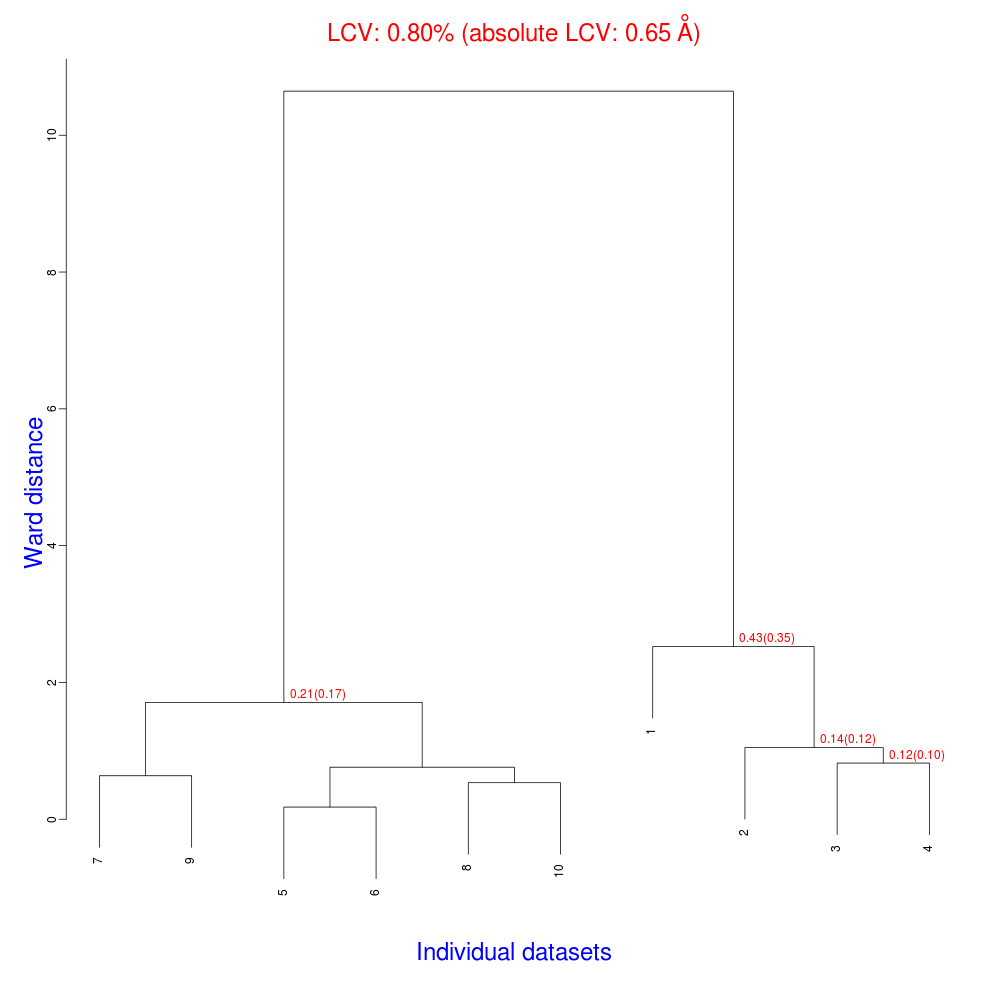
As numbering of the 10 data sets has not been modified, one can see that clusters for these data include the same elements of corresponding clusters in the previous example, considering that data sets numbered 11 to 14 have been excluded from the analysis.
Having shown how BLEND in analysis mode can be re-executed using a subset of data, let us go back to the previous working directory selecting “BLEND_TUTORIAL_INSULIN” project directory in ccp4i. Let's try and increase or decrease image cutting due to radiation damage, by using the RADFRAC keyword. An input value of 0.85 means that we would like to keep only images for which intensities have, on average, been dampened by less than 15% of their initial value. Input for RADFRAC can be included via ccp4i as shown here:
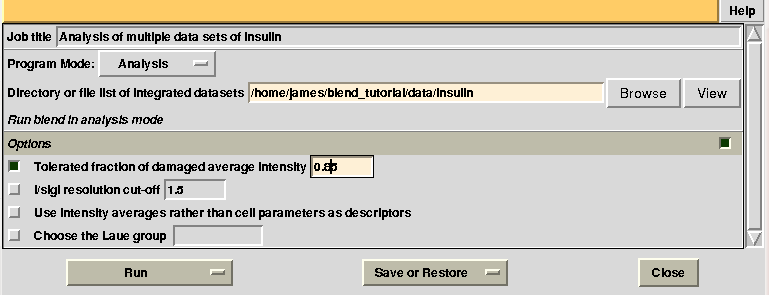
This time “FINAL_list_of_files.dat” shows some difference in the number of images discarded. For data set 9 discarded images range 102 to 122, while before it ranged 108 to 122, and images 93 to 100 are now discarded from data set 7, while no images were previously discarded. A small change also occurs for data set 1:

Let's modify RADFRAC once more and use 0.65, rather than 0.85. As a result:
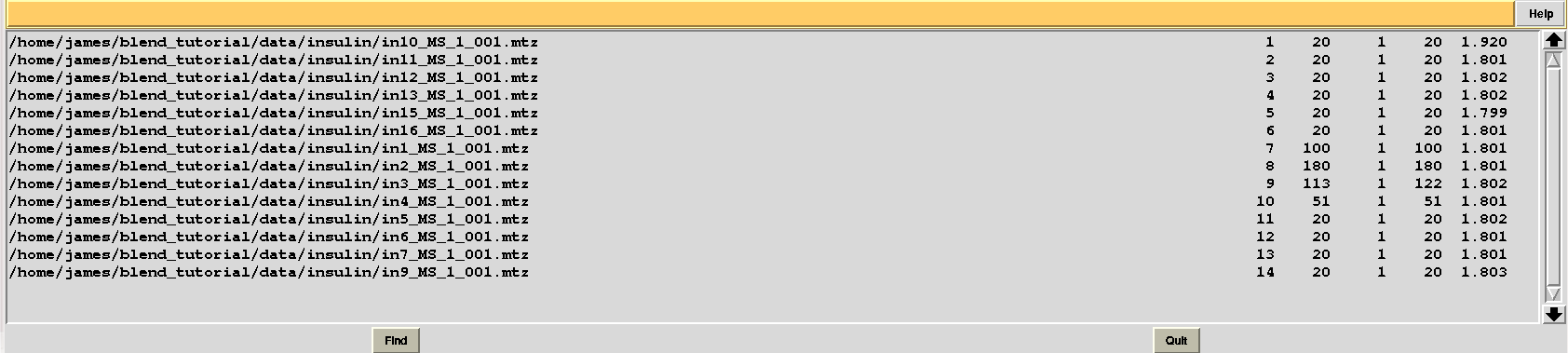
Images discarded are now less because we have allowed for intensities to be dampened, on average, by up to 35% of their starting value. A similar data exclusion can be applied in resolution terms, by changing value to the keyword ISIGI. For instance, let's re-run BLEND in analysis mode (with RADFRAC 0.75 - default value as the box "Tolerated fraction of damaged average intensity" is not selected), fixing ISIGI to 3:
Results should be produced in directory “blend_4”. The “FINAL_list_of_files.dat” looks like the following:
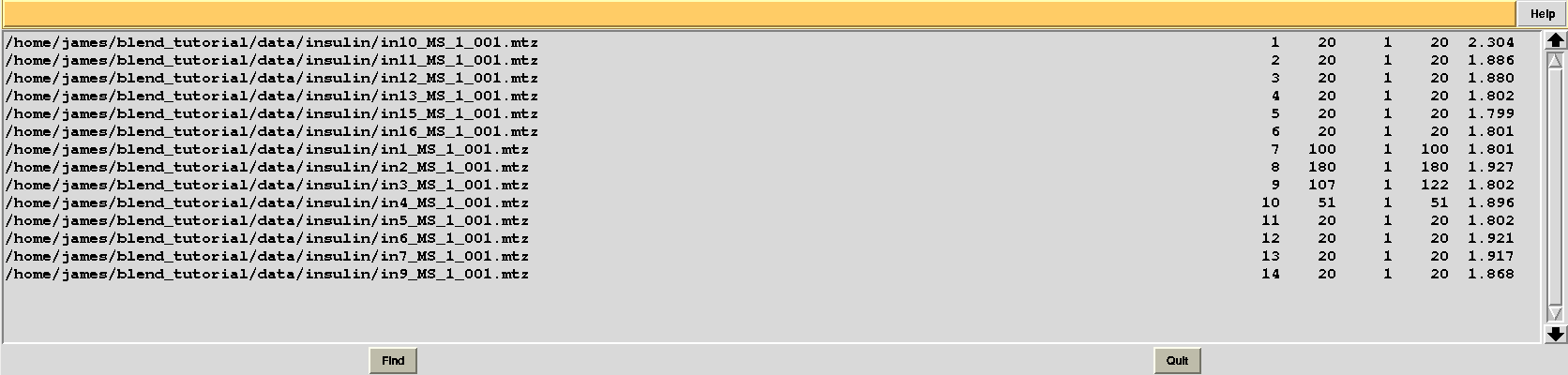
Resolution cutoffs have now changed. For the first data set the highest is 2.304, while previously it was 1.920. We expect an increase if the value of ISIGI is lowered. This is normally the case. But in the specific insulin example data are exceptionally good, and their signal-to-noise ratio is quite high to the very highest resolution. Thus, results will not change here if values lower than 1.5 are used for ISIGI.
Let's pay a closer look at the results of cluster analysis, as described in file “CLUSTERS.txt”:

Cluster numbers and corresponding data sets are listed in column 1 and column 6 respectively. The second column reports how many data sets form the specific cluster, while in column 3 it is found the exact height at which data sets merge to form the cluster. Columns 4 and 5 lists all LCV and aLCV values. The dendrogram encourages people to look for grouping of individual data sets. Larger groups attract one's attention more than smaller groups. In this sense the presence of two clusters, cluster (1 2 3 4 14) and cluster (5 6 7 8 9 10 11 12 13), is immediately perceived. Very soon, though, we realise that the second cluster is made up of two smaller clusters, cluster (5 6 8 10) and cluster (7 9 11 12 13). Finer details and even smaller clusters can be singled out by proceeding to the lower part of the dendrogram. This way of proceeding through the dendrogram interpretation can have some validity, but is highly subjective. In BLEND it is found that a more logical, straightforward and objective way of interpreting the dendrogram is to assume that each node can represent a valid data set. Subsequent operations of space group assignment (POINTLESS) and scaling (AIMLESS) determine the degree of validity. This interpretation is accomplished with BLEND synthesis execution mode. From “CLUSTERS.txt” we read that everything merges at height 13.670. If the user wishes to produce results for all nodes under a certain height, it will suffice to specify this in the GUI. For instance, to produce results for all nodes in the insulin example, the BLEND section of the GUI will look like this:
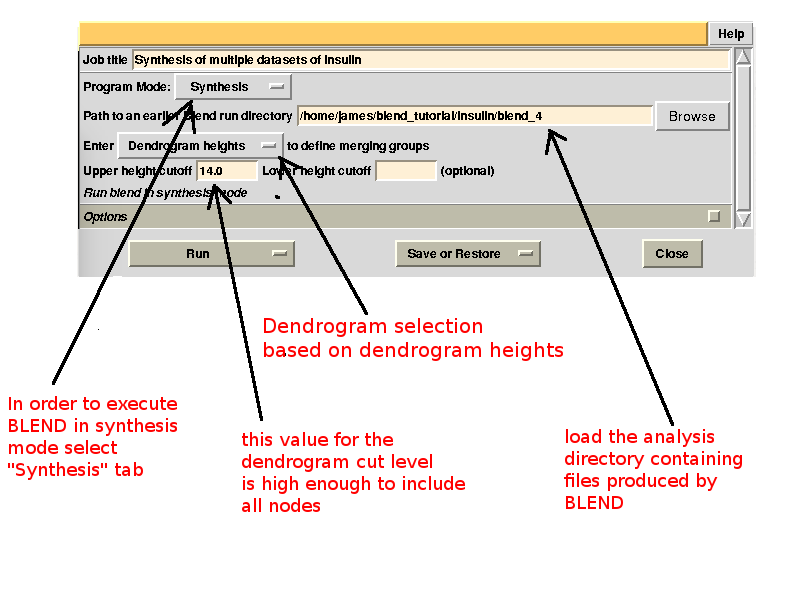
Cluster selection can also be based on LCV or aLCV values. This might make sense when one's only interest is in finding the most compact groups of crystals. The selection of all clusters just reported can be also achieved by selecting "aLCV values" instead of "Dendrogram heights" and inputting for example "0.85" (so to cover all clusters), rather than "14".
After execution, several files will be produced in directory “blend_5” (this is the fifth job for the GUI under project directory “BLEND_TUTORIAL_INSULIN”), including another directory packed with files, called “merged_files”:
In this directory one can find unscaled (produced by POINTLESS) and scaled (produced by AIMLESS) files corresponding to all nodes selected in the dendrogram. Final overall merging statistics can be found in the “MERGING_STATISTICS.info” file:
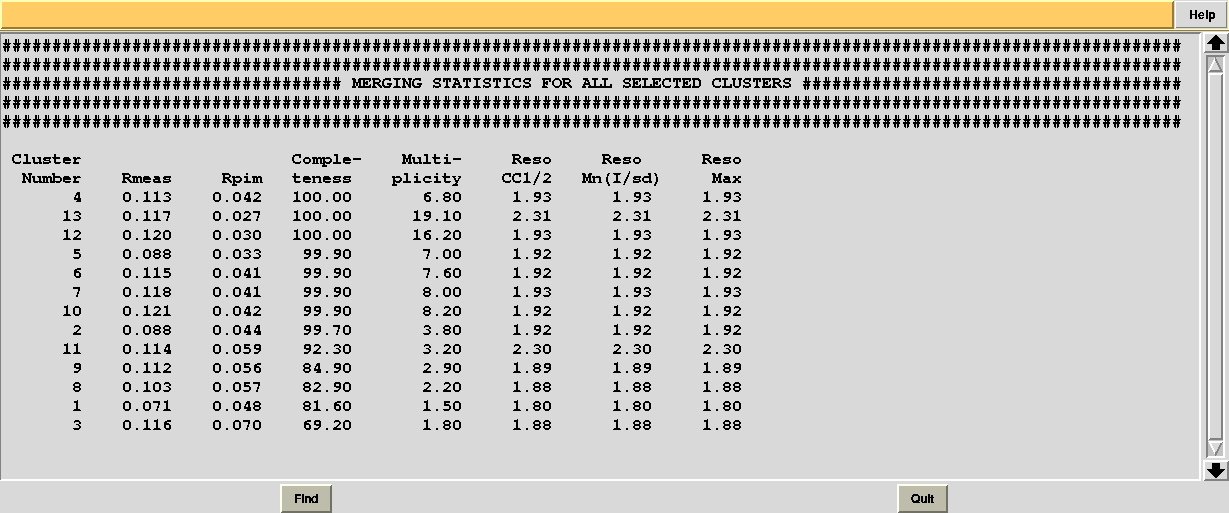
Values are sorted on increasing completeness, followed by lower Rmeas. Higher completeness is often associated with high redundancy (multiplicity), but also with poor isomorphism, because many more data sets are needed for both completeness and multiplicity to increase. For example, cluster 12 is 100% complete, as it is cluster 4, but this last cluster has lower Rmeas, quite certainly because it is the union of just two data sets, while cluster 12 includes nine data sets. A good result is represented by cluster 2, which has Rmeas = 0.088 and it is 99.7% complete. Cluster 5 is an even better choice; it has same Rmeas than cluster 2, but a much lower Rpim. It is a nearly complete set and has high multiplicity. This is what should probably be used. A visual version of the above table is provided by the “Rmeas_vs_Cmpl.png” or “Rmeas_vs_Cmpl.ps” plot:
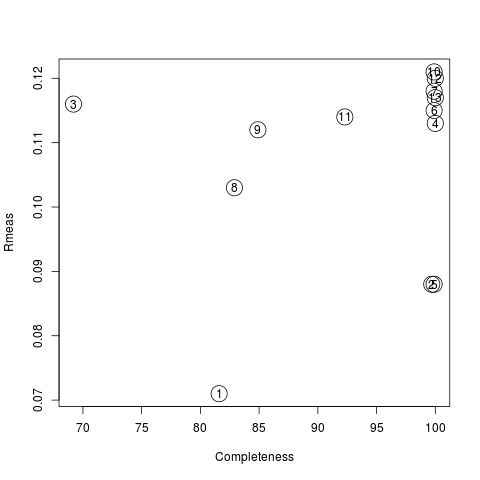
From the overall view provided by this plot we can see that 2 and 5 are the best data sets in terms of statistics and completeness (they are barely visible because they overlap on top of each other). Unscaled and scaled files related to all results, in the “merged_files” directory, can be inspected with the usual CCP4 tools (try, for instance, to view “scaled_005.mtz”). POINTLESS and AIMLESS logs for all unscaled and scaled files are also included in this directory. For instance, the final part of the AIMLESS log file for cluster 5 shows:
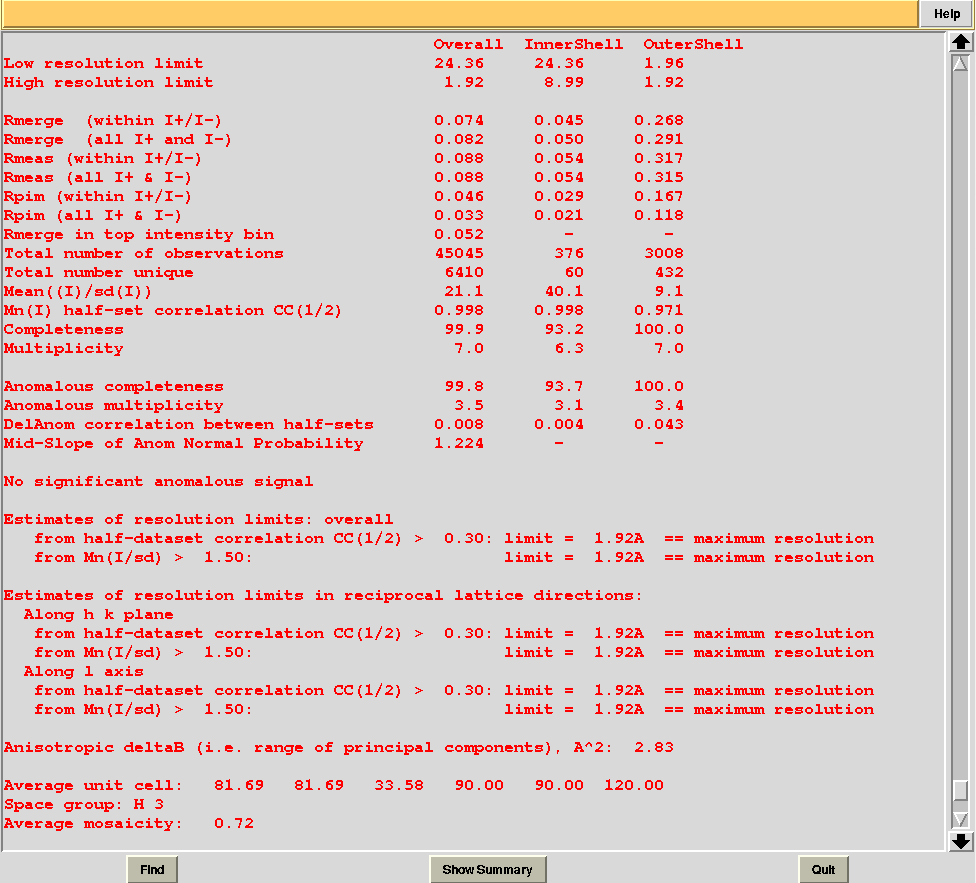
Overall merging statistics can also be explored selecting "View Files from Job" → "View Job Results (new style)":
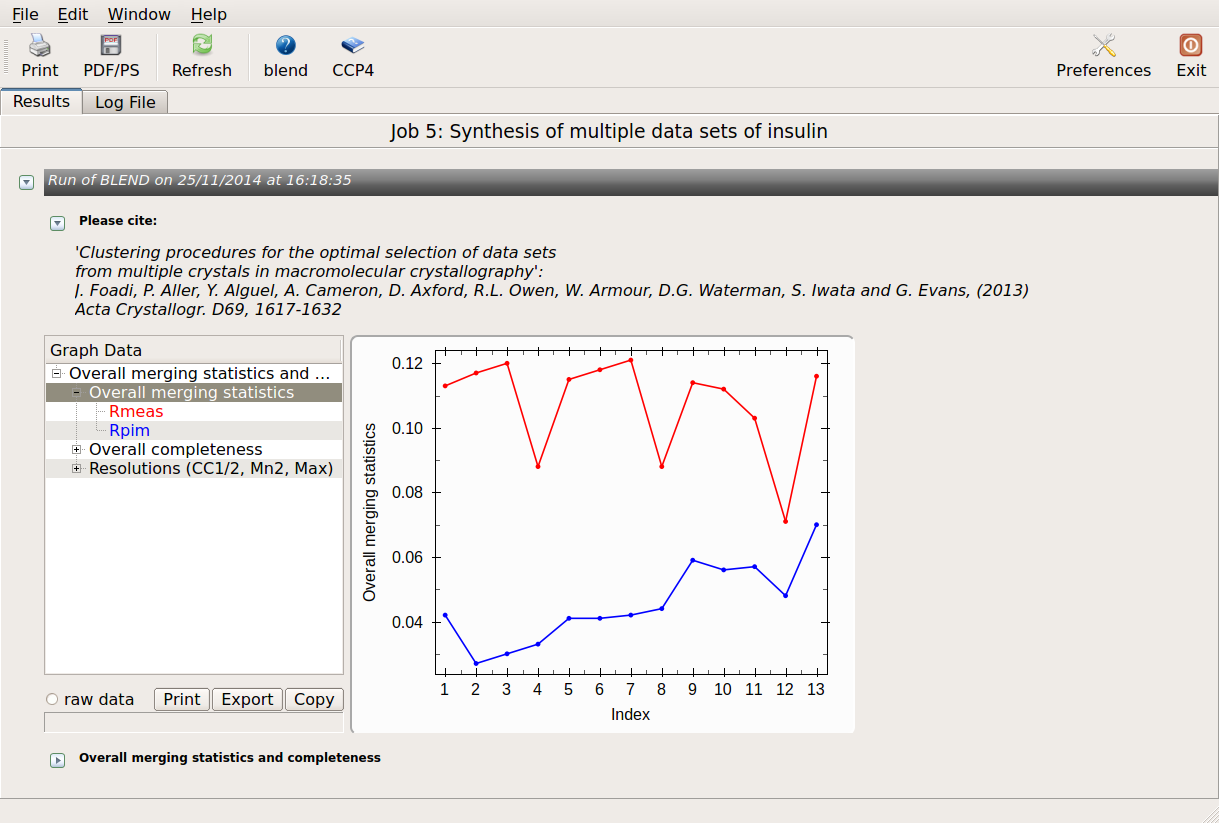
There might be reasons for which the user wishes to calculate results for certain nodes only, rather than for all nodes of the dendrogram. This can be done using two numerical values, rather than one, in BLEND synthesis. Suppose we only want to calculate data for cluster (5 6 8 10). This cluster merges at a height of 0.744 (check “CLUSTERS.txt”). Two numerical values in between which 0.744 and no other nodes is included are 0.745 and 0.743. Thus, to produce a synthesis just for this cluster, one can run BLEND as follows:
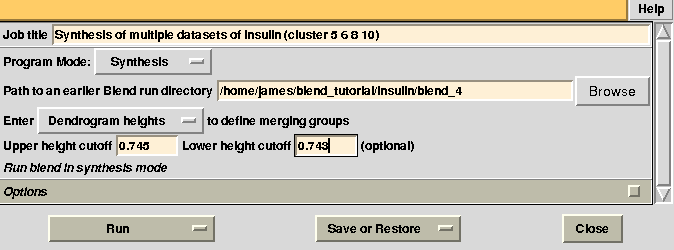
Statistics and related files for only one data set are contained within the “merged_files” directory. This job could have been executed more easily in combination mode, simply selecting the specific cluster in the graphical interface (see later). Suppose we want to extend resolution of the previous result from 1.93 to 1.80. BLEND can be again executed in synthesis mode using the appropriate "RESOLUTION" keyword:
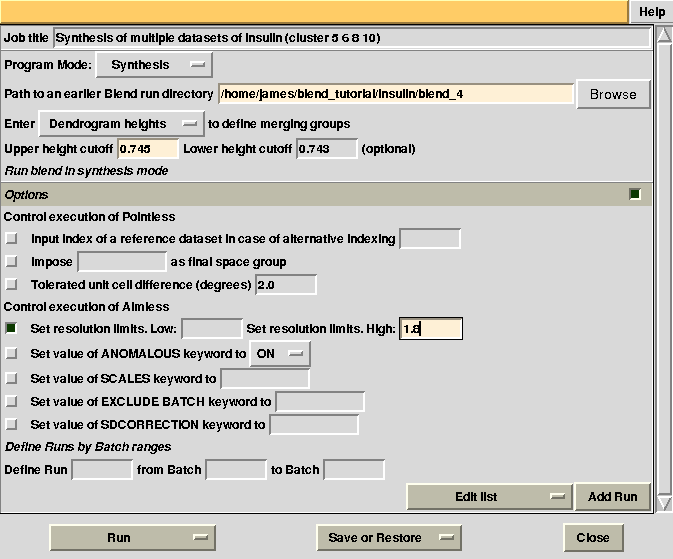
Check statistics under the new “merged_files” directory, they should have changed to include the desired resolution cutoff. "MERGING_STATISTICS.info" allows people to gain a better understanding of the ability individual data sets have to form larger, more complete and isomorphous groups. For example, in job number 5, it clearly emerges that cluster 5, composed of data sets 7, 9, 13, has good merging statistics and it is very complete. The two additional data sets joining 7, 9, 13 and forming cluster 10, i.e. data sets 11 and 12, make the situation worse, in terms of merging statistics.
It is possible to find out which data sets make the statistics worse by running BLEND in combination mode. For example, we might want to investigate whether data set 11 or data set 12 make statistics for cluster 5 worse. First we try with 11. The group formed by data sets 7, 9, 11 and 13 is equivalent to cluster 10 without data set 12. This can be specified in the graphical interface, to save typing time (this option becomes very useful when handling several data sets). As we would like to compare results with what has been computed for cluster 10 in BLEND job 5, we also fix the highest resolution to the same value used in that run, 1.92 Å:
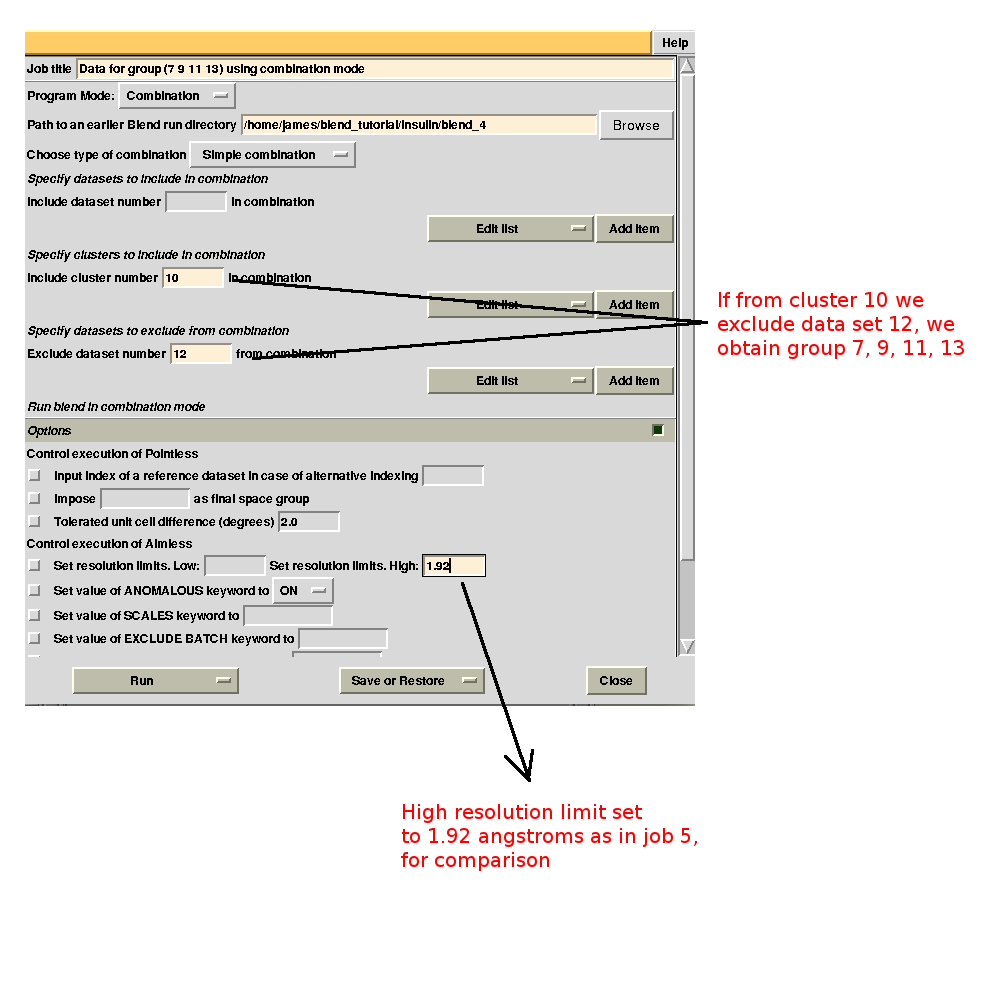
Merging statistics for this specific combination can be found in the sub-directory "combined_files" of directory "blend_8" (as Job 8 was the BLEND job just executed). Next we try with 12. If the "earlier directory" from which the new BLEND job will be executed coincides with the output directory of Job 8, all results will be collated in a same directory. This becomes very useful when many different groups are to be investigated. The interface for the new job is, thus:
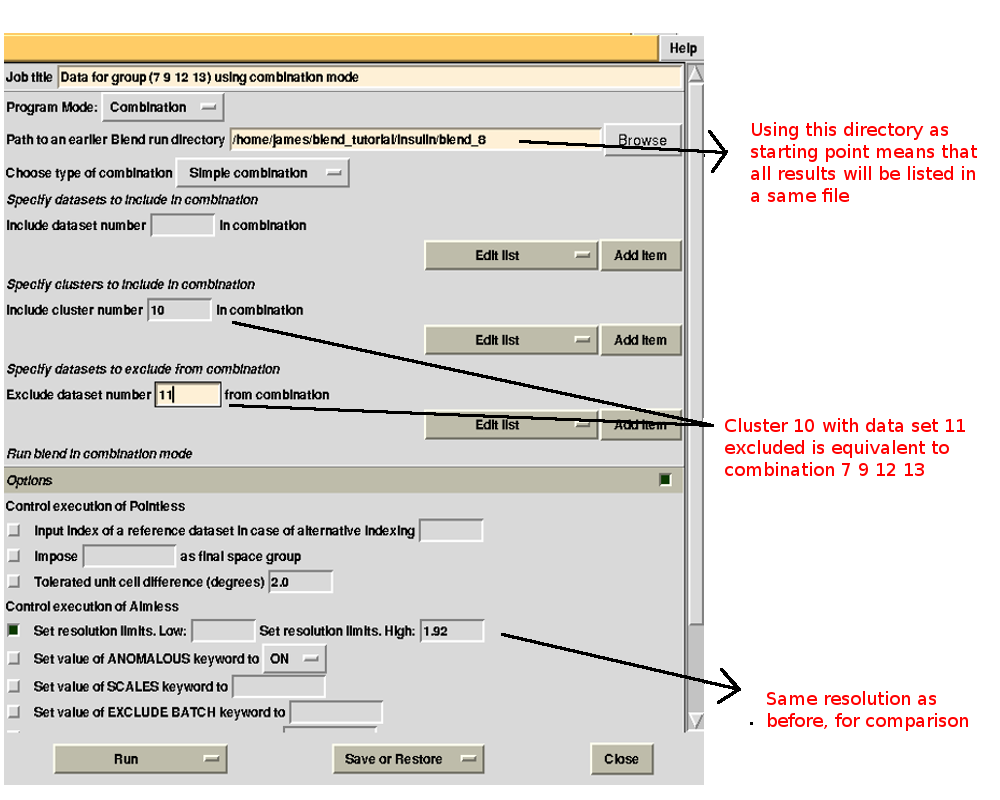
Checking file "MERGING_STATISTICS.info" in sub-directory "combined_files" of directory "blend_9" leads to the conclusion that statistics are better when using data set 11 (group 1 because we ran combination job for group 7, 9, 11, 13 first):
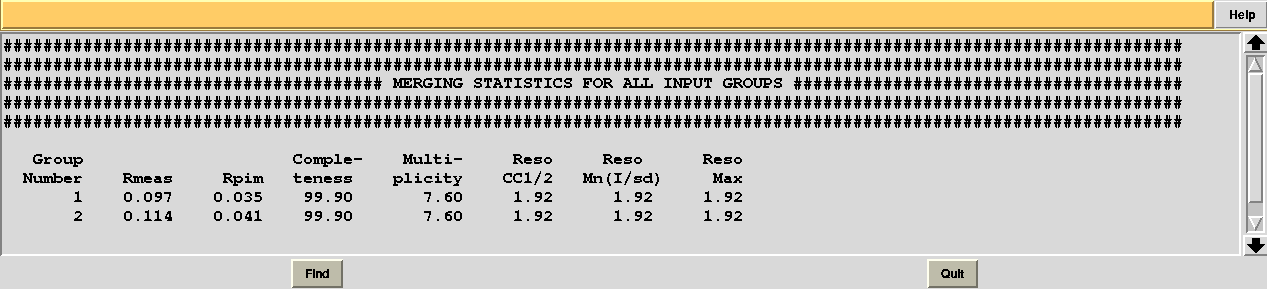
Thus we can decide to "filter out" data set 12 from cluster 10, in order to achieve a better composite data set.
b) Without CCP4I
Like any other CCP4 program, BLEND can be executed without using ccp4i, simply typing commands at the keyboard in an interactive command shell. This, quite often, makes the program more flexible. Here we will briefly repeat all steps previously carried out with ccp4i. First a new directory, named for example "insulin_no_ccp4i" will be created in order to keep this part of the tutorial separate from the rest:
mkdir insulin_no_ccp4
cd insulin_no_ccp4
Next we run BLEND in analysis mode using all 14 files in the "data/insulin" directory:
blend -a ../data/insulin
After start the program immediately halts, waiting for keywords input; in this specific instance we simply push the "Enter" key once, thus forcing BLEND to use default values. At completion the program produces the following files:
BLEND.RData
BLEND_SUMMARY.txt
CLUSTERS.txt
FINAL_list_of_files.dat
mtz_names.dat
tree.png
tree.ps
whose meaning and content has been previously explained. To show how "mtz_names.dat" can be re-used for a different run of BLEND in analysis mode, and similarly to what done previously in this tutorial, let's create a directory called "amended_insulin" and copy "mtz_names.dat" in it:
mkdir amended_insulin
cd amended_insulin
cp ../mtz_names.dat original.dat
Then "original.dat" is edited to remove the last four lines, thus creating a different input, consisting only of 10 out of 14 data sets. Next, the program is executed using "original.dat" as input:
blend -a original.dat
Results of this run are similar to what described before. To carry on let's move back one level to "insulin_no_ccp4i":
cd ../
If a log file is needed, then a keyword file has to be prepared before execution. Let's call this file "bkeys.dat", and play both with RADFRAC and ISIGI keywords, changing from their default values 0.75 and 1.5 to the new values 0.65 and 3.0. The file looks like this:
RADFRAC 0.65
ISIGI 3.0
Then BLEND is executed again using "mtz_names.dat" as input:
blend -a mtz_names.dat < bkeys.dat > blend_analysis.log
After completion, "blend_analysis.log" can be either read using any editor, or viewed graphically with the CCP4 program "logview":
logview blend_analysis.log
To run BLEND in synthesis mode and produce scaled data for all nodes of the dendrogram, we could type:
blend -s 14 < bkeys.dat > blend_synthesis.log
But, just to show that the same results are obtained using aLCV, rather than the dendrogram's height, we executed BLEND in the following way:
blend -saLCV 0.85 < bkeys.dat > blend_synthesis.log
At completion the result can, again, be viewed using "logview":
logview blend_synthesis.log
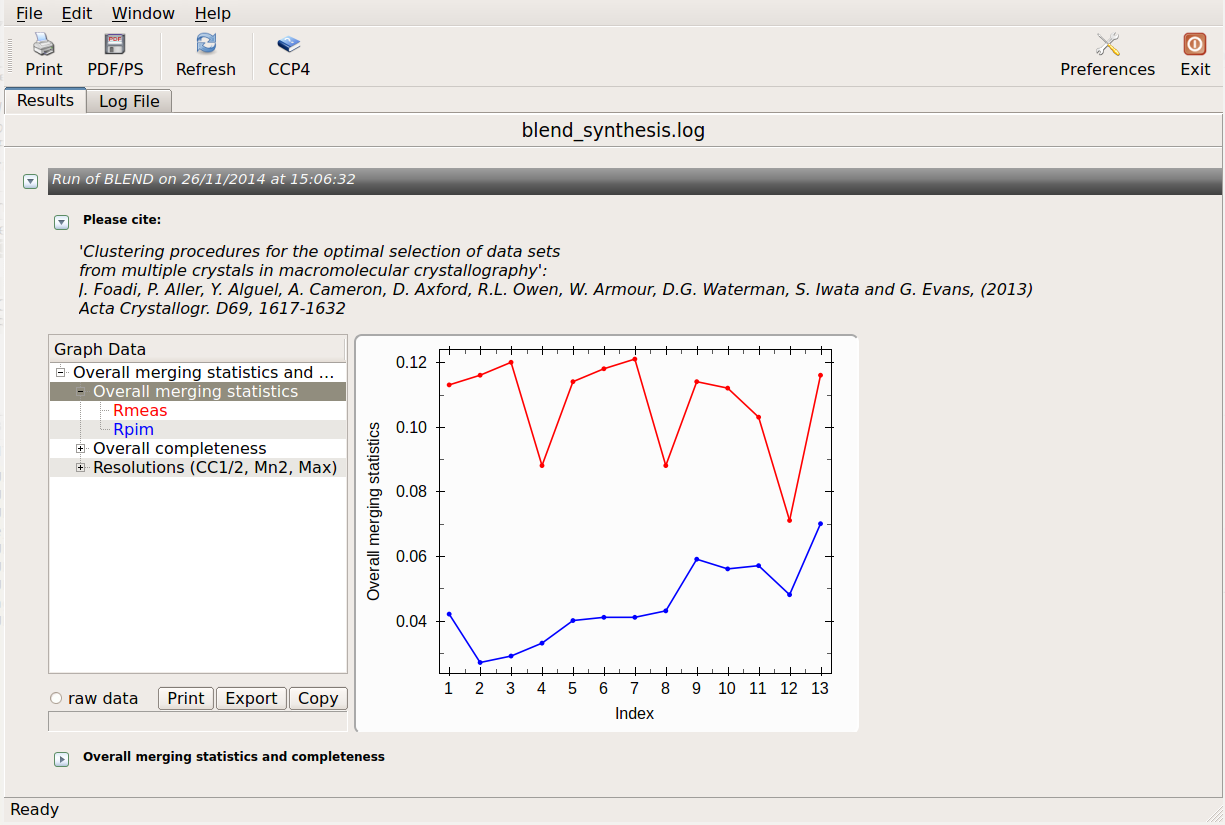
All unscaled and scaled mtz files, and all POINTLESS and AIMLESS logs, are contained in the "merged_files" directory. Logs can be viewed using "logview" or "loggraph". For example:
loggraph merged_files/aimless_005.log
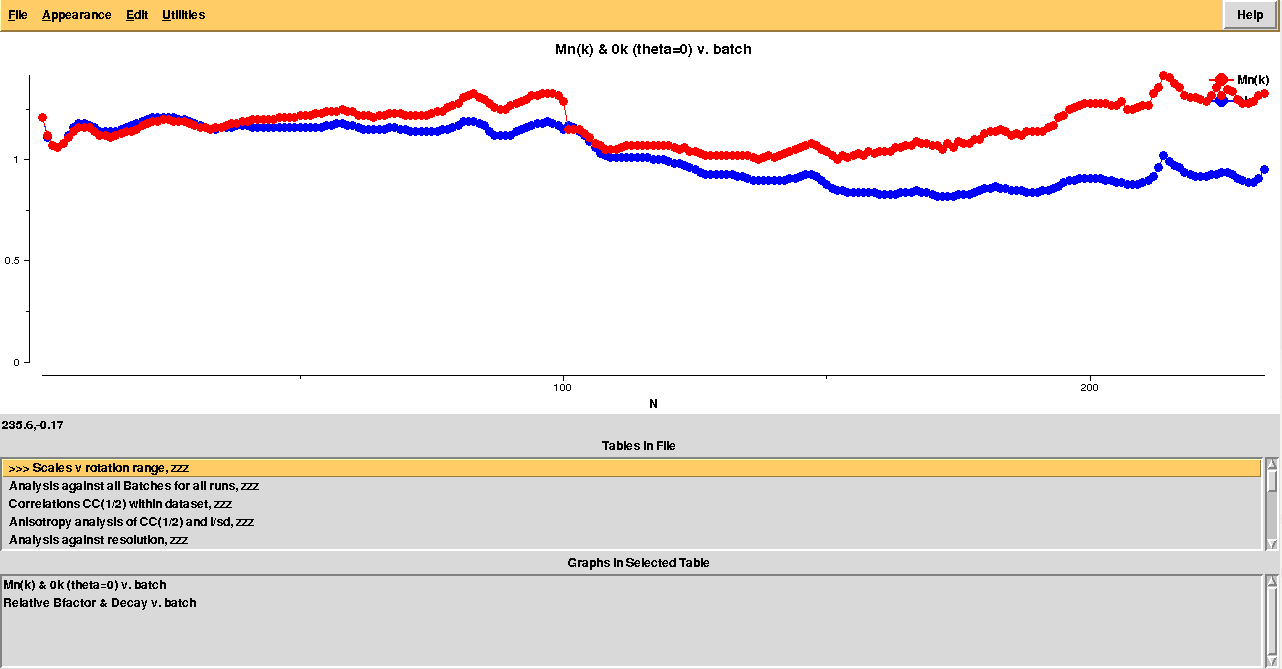
In order to execute BLEND in combination mode, so to scale groups of data sets not corresponding to any cluster, it is handy to copy and paste numeric strings from the sixth column in file "CLUSTERS.txt". Another, perhaps quicker, possibility is to use the special syntax created to summarize groups of data sets (see documentation).
For instance, the group of data sets (7, 8, 9, 10, 13) is simply the union of clusters 4 and 5. This can be represented with the two cluster numbers within square brackets, "[4,5]". The command line to run BLEND in combination mode over this group would be, in this case (using same keywords file as before):
blend -c [4,5] < bkeys.dat
Or, group (7, 9, 11, 13) is essentially cluster 10 with the exclusion of data set 12. This can be summarized with the following string, "[10] [[12]]", where the double-squared brackets include all data sets to be excluded during execution:
blend -c [10] [[12]] < bkeys.dat
Many more runs in combination mode can be executed within the same directory. All files produced can be found in directory "combined_files".
How to obtain complete data sets, while preserving isomorphism with BLEND
These data come from hen egg lysozyme with two different procedures. The first group of 12 10-degree sweeps has been collected with pure lysozyme crystals at room temperature from crystallisation plates. The second group of 16 10-degree sweeps has been obtained from crystals soaked in a solution of sodium bromide; data have been collected at the Diamond Light Source synchrotron with a wavelength close to the bromide peak. Data collection went further than 10 degrees for each data set, but we have limited sweeps in order to replicate similar situations met when radiation damage acts fast. For this tutorial in situ data have been stacked with data collected at low temperature and from a slightly different structure (additional bromide content) in order to demonstrate the interplay between completeness and non-isomorphism.
Let's create a new directory, called "lysozyme", in the "blend_tutorial" directory, and associate the new project directory "BLEND_TUTORIAL_LYSOZYME" to "lysozyme":
mkdir lysozyme
Next, we run BLEND in analysis mode using all mtz files in "data/lysozyme":
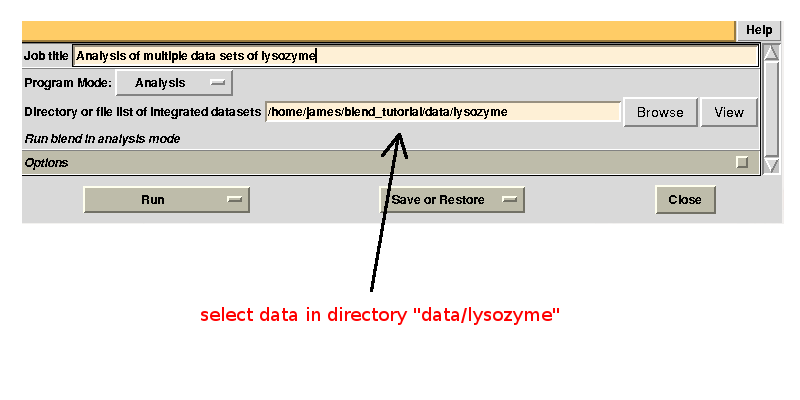
In the group of files produced by this run the following dendrogram (file "tree.ps" or "tree.png") is found:
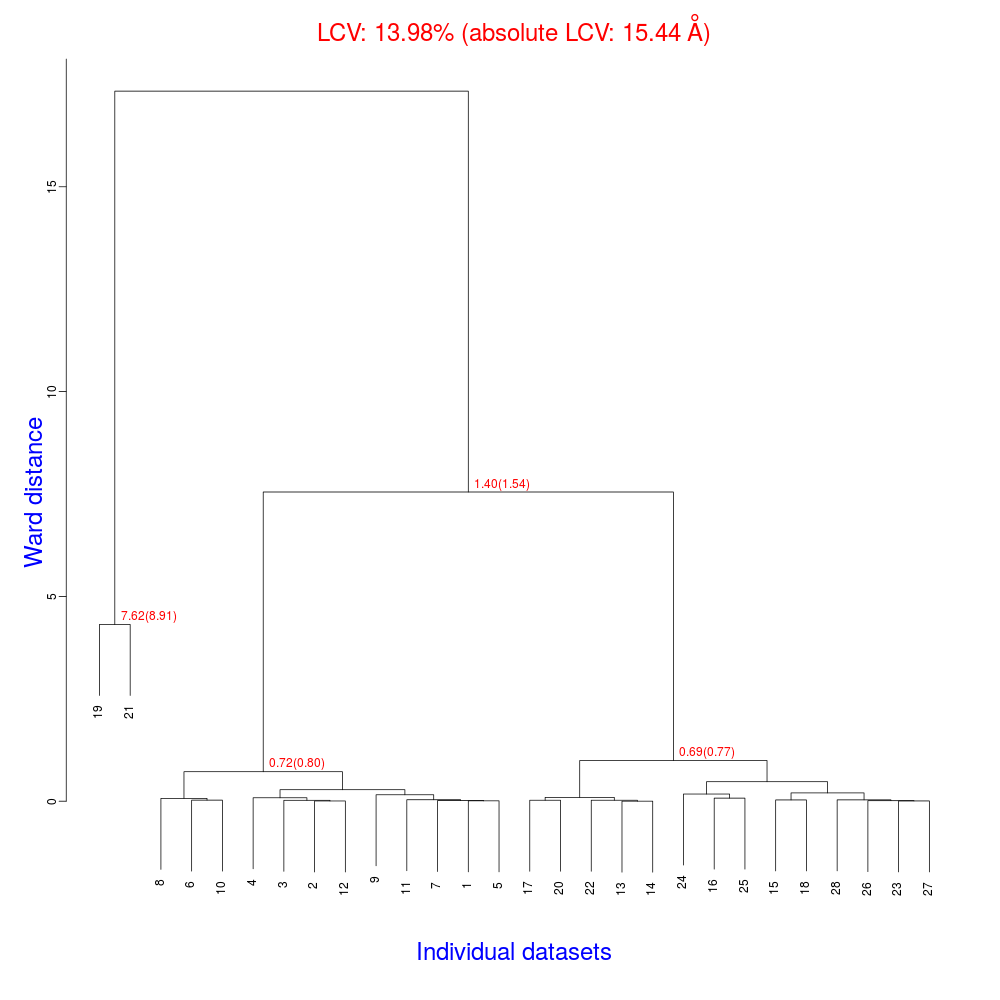
The overall Linear Cell Variation (LCV) has a high value. This is certainly caused by the two of crystals in the isolated branch on the left of the dendrogram, as it can be confirmed by looking at cell parameters plots in BLEND log file (new style):
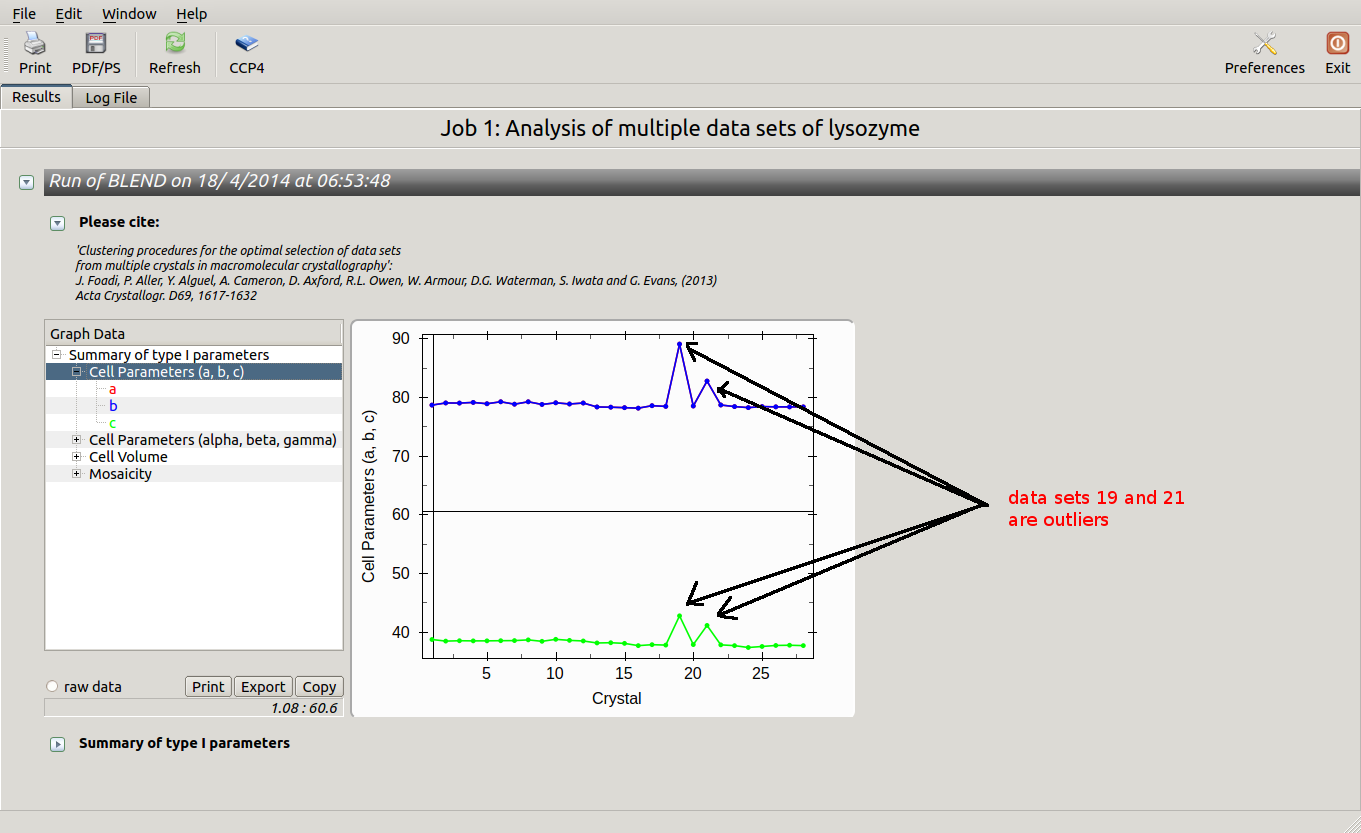
Data consist of many short sweeps (not useful to form a complete data set), therefore we can get rid of sweeps 19 and 21 without loosing much data and re-run BLEND in analysis mode. One way of executing BLEND using the same data without sweeps 19 and 21 is by copying file "mtz_names.dat" into "original.dat" and deleting the lines pointing to these sweeps:
cp lysozyme/blend_1/mtz_names.dat lysozyme/original.dat
[edit "lysozyme/original.dat" and delete lines containing "dataset_019.mtz" and "dataset_021.mtz"]
BLEND is now executed using "original.dat" as input:
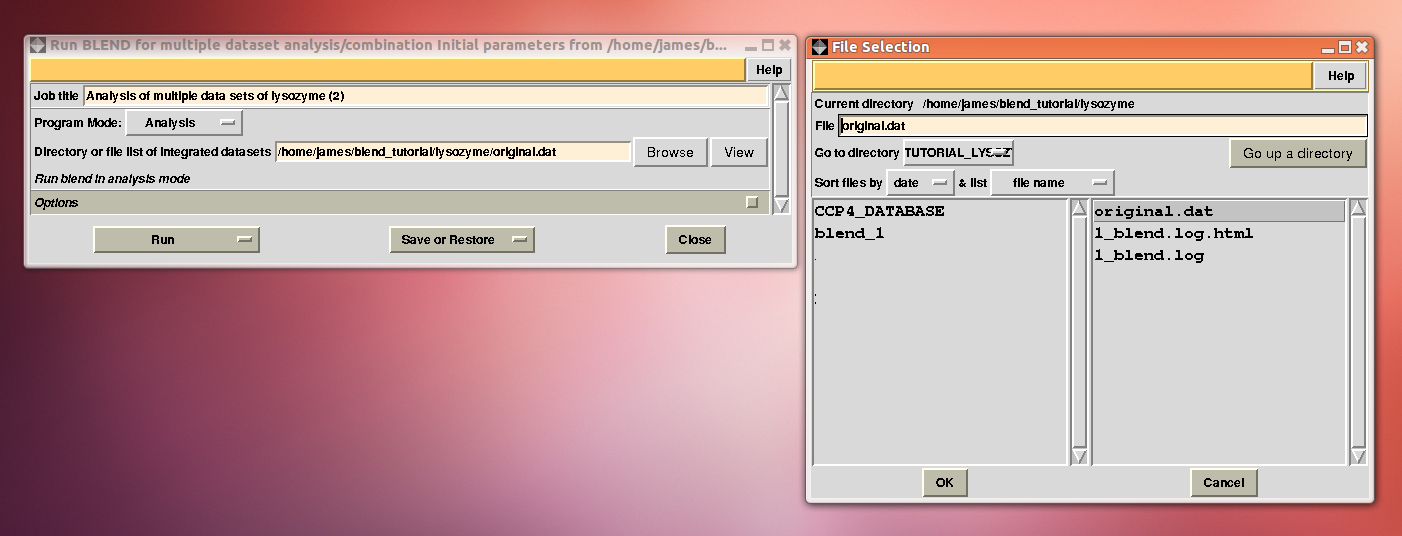
The ensuing dendrogram is:
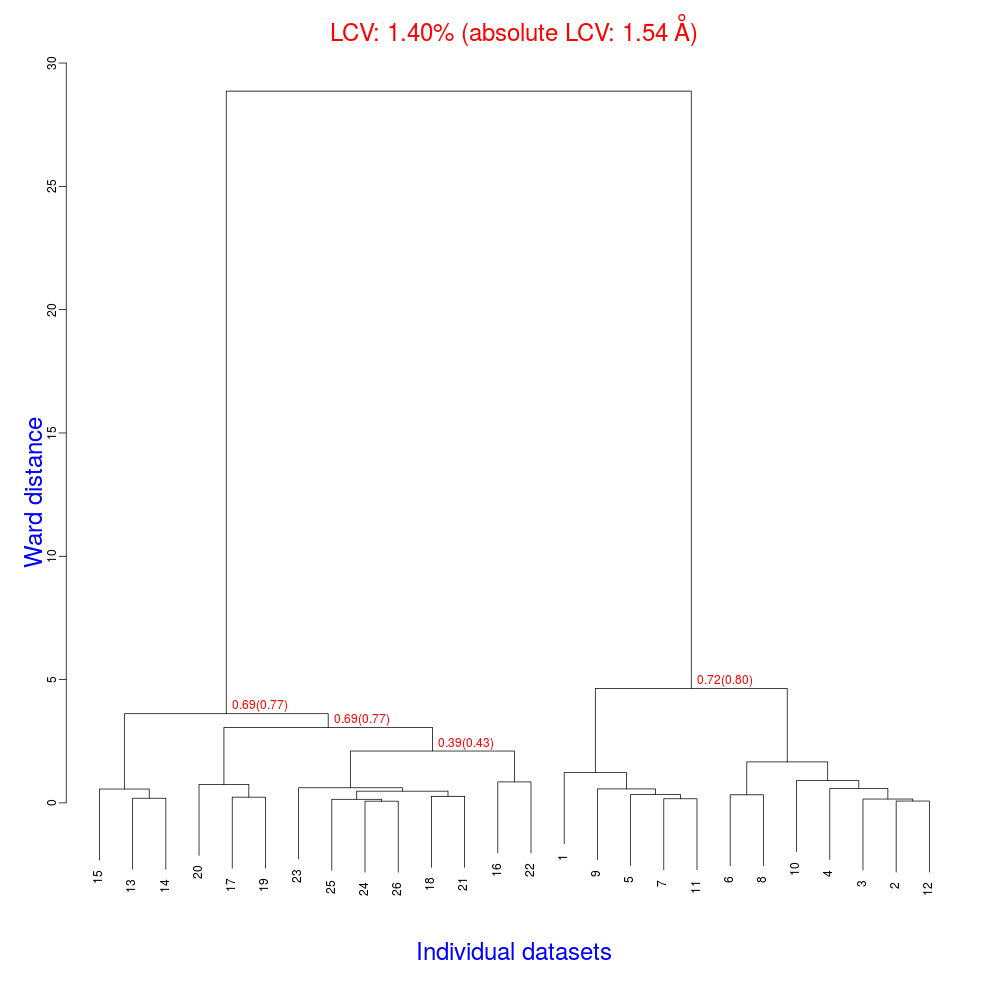
As we can see, the new LCV is much smaller, although the value 1.40% (and the absolute LCV value 1.54 Å) still indicates some non-isomorphism among all crystals. We know this is the case because crystals have been assembled from differently-prepared structures. Indeed, LCV values computed separately for the left branch and for the right branch of the dendrogram return 0.69% and 0.72%, respectively. Incidentally, data sets group 1 – 12 has different features from group 13 – 26 (remember, two sweeps missing from the initial group!) because it corresponds to a different collection from a different group of crystals. This coarse division, observed in the two separate dendrogram's branches, can also be detected by looking at the different mosaicity trends in BLEND log file:
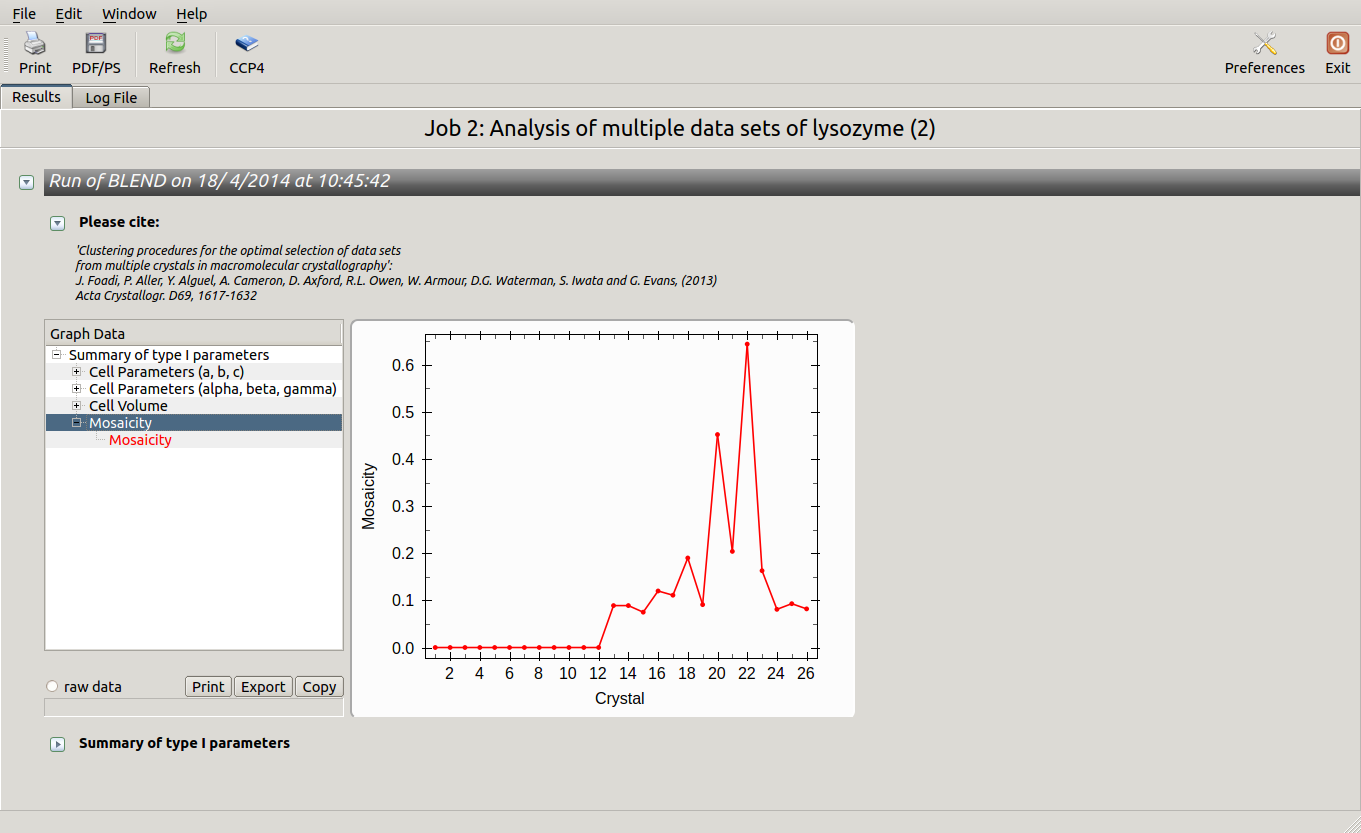
Scaled data for all nodes of the new dendrogram can be computed running BLEND in synthesis mode. We know that all data should belong to space group P 43 21 2. We also would like to use the same resolution for all data, say 1.9 Å. Accordingly, the program is executed using the following keywords:
CHOOSE SPACEGROUP P 43 21 2
TOLERANCE 100
RESO HIGH 1.9
Keyword TOLERANCE is set to a high value to bypass the default tendency of POINTLESS to stop execution when cell parameters differ too much. Considering now the dendrogram, using only one value for the height means that the program would process all nodes below this value. In the "CLUSTERS.txt" file it can be seen that these have values smaller or equal to 28.861; thus any value larger than this produces the desired scaled data:
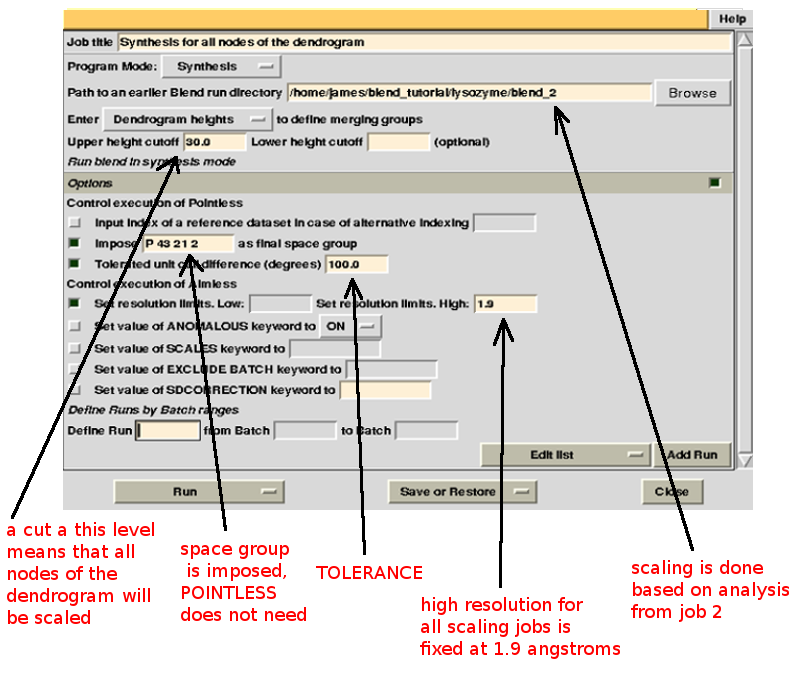
This run will take a while to complete (many clusters!). At the end of execution a new "merged_files" directory is created with scaled files and merging statistics tabulated. Here is a view of the final statistics in BLEND log:
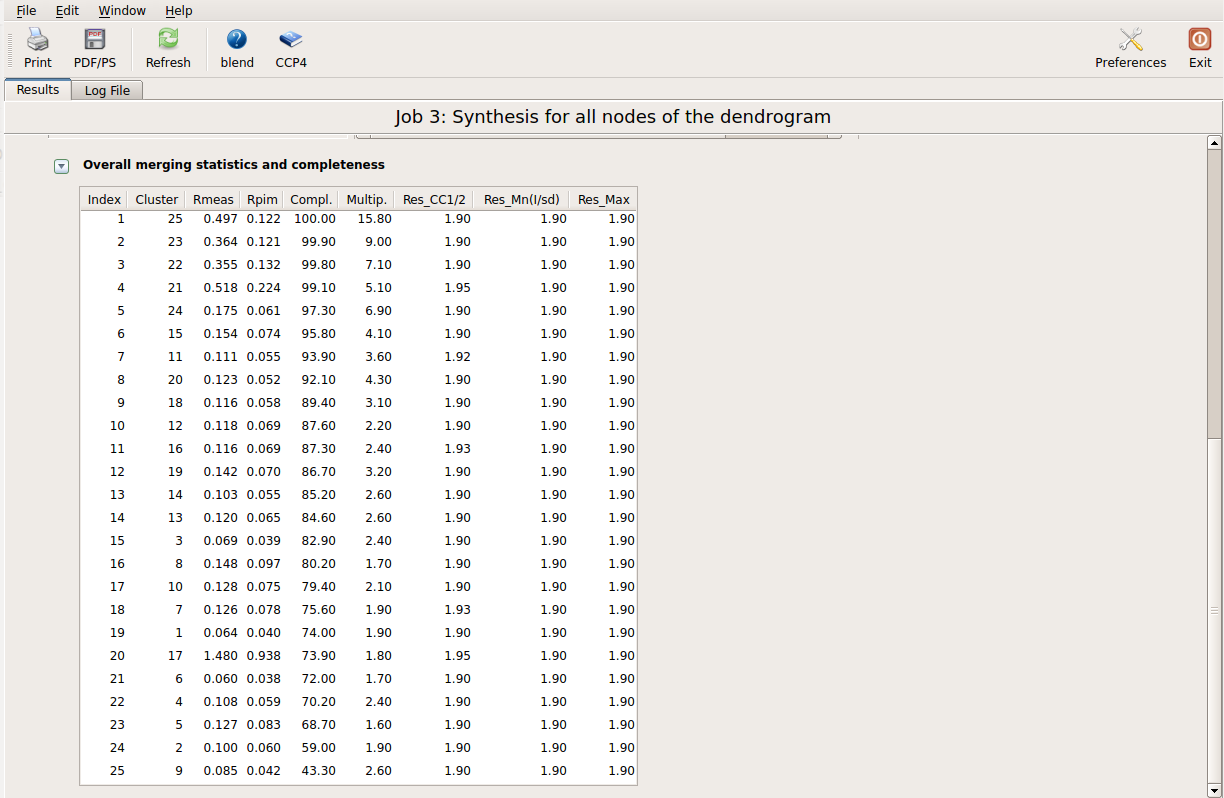
Scaled files corresponding to these results are included in the "merged_files" directory, and they are ready for further investigations. Some of the statistics can be improved by filtering out certain sweeps from specific clusters. A visualization of the above table can be conveniently created running BLEND in graphics mode. The program needs to be pointed at the directory in which the run in synthesis mode has been executed. Three types of results from the statistics produced by AIMLESS will be annotated in the dendrogram around a user-specified cluster. Furthermore details of the dendrogram are determined by the user-selected number of levels down the chosen cluster. For the specific case treated here we would like to view merging statistics in an annotated dendrogram starting from the top cluster (cluster 25) and going down 10 levels (this is normally enough for a first view of the tree). For executing this mode the gui is arranged as in the following picture:
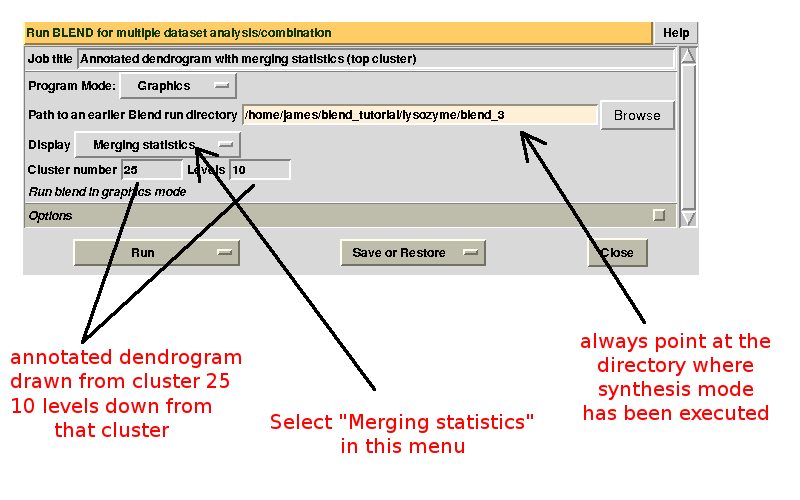
And this is the annotated dendrogram created:
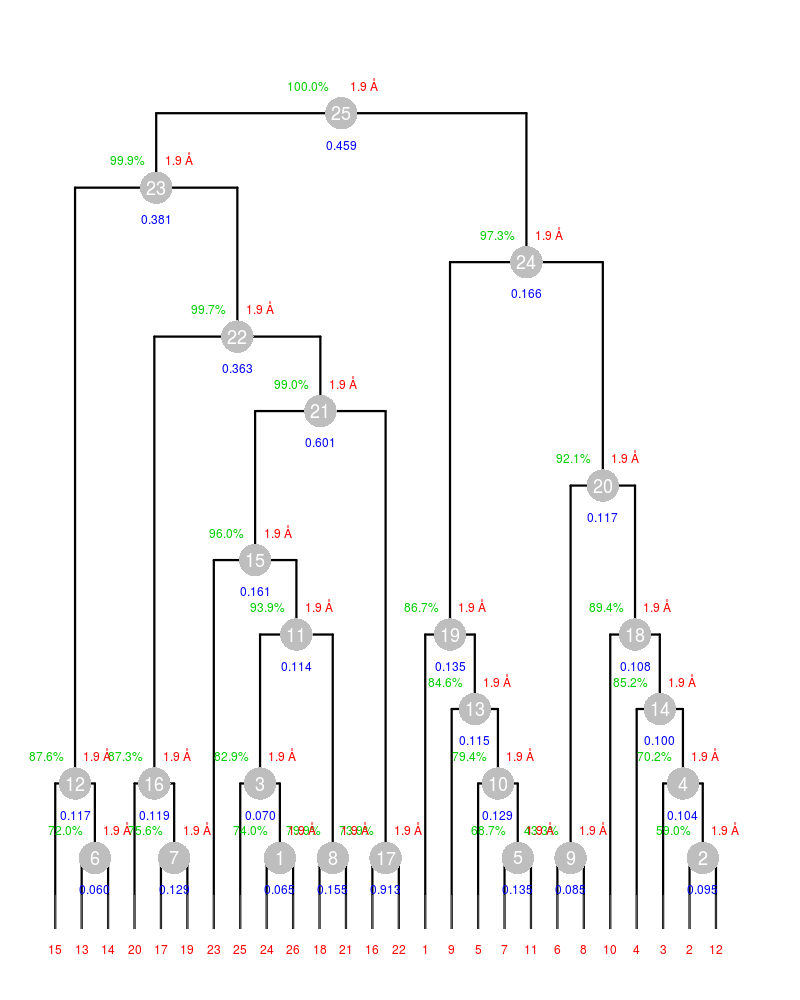
In the above annotated dendrogram clusters at a same level are formed by the same number of data sets. If the dendrogram is densely populated it might be difficult to see which data set belong to which cluster. For example clusters 13 and 14 are both formed by 4 data sets; clusters 18 and 19 both formed by 5 data sets. Clusters 21, 22, 23 and 25 have high Rmeas values, while values for clusters 15 and 24 can, perhaps, be improved. The main way to improve statistics at this point is by running BLEND in combination mode. Let's try, for instance, to improve the statistics for cluster 15. As the Rmeas value jumps from 0.114 for cluster 11 to 0.161 for cluster 15 and as cluster 15 is simply cluster 11 with the addition of sweep 23, it is straightforward to impute the reason for the increase to the bad quality of sweep 23. But there might also be other possibilities. For example sweep 23 does not match well some of the sweeps composing cluster 11; by excluding these sweeps, then, statistics might improve. In conclusion we can try all combinations of data sets forming cluster 15, excluding one data set at a time. For all combinations after the first we will have to make sure that the path to an earlier BLEND directory points to the results directory of the previous combination. So for the first combination:
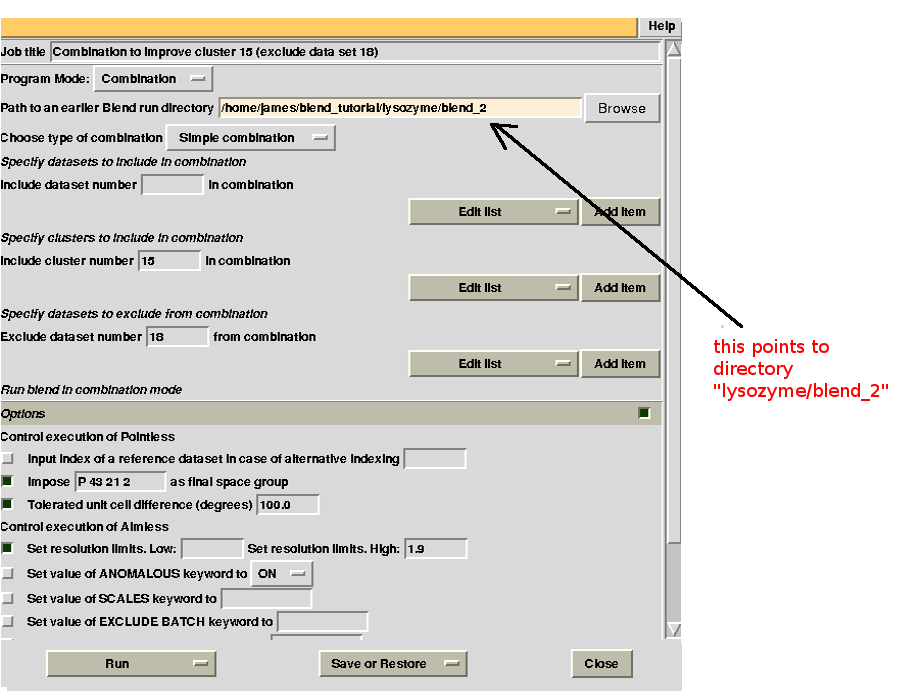
we have excluded data set 18. For the second combination:
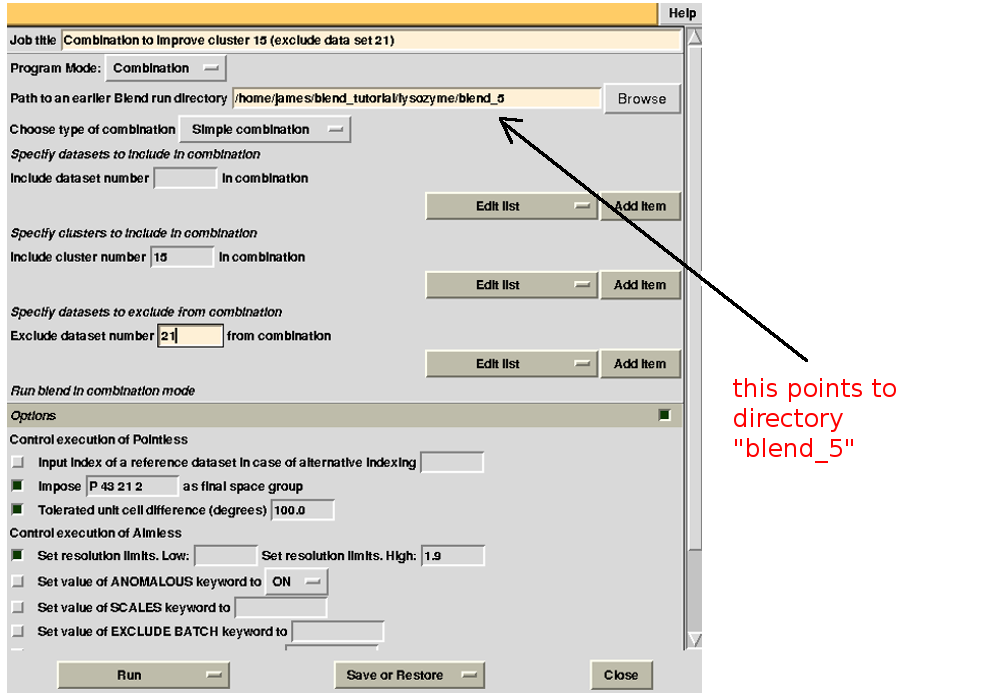
To summarize, for all combinations tried to improve statistics for cluster 15, we have:
combination 21 23 24 25 26 (equivalent to cluster 15 without data set 18) points to directory “lysozyme/blend_2”
combination 18 23 24 25 26 (equivalent to cluster 15 without data set 21) points to directory “lysozyme/blend_5”
combination 18 21 24 25 26 (equivalent to cluster 15 without data set 23) points to directory “lysozyme/blend_6”
combination 18 21 23 25 26 (equivalent to cluster 15 without data set 24) points to directory “lysozyme/blend_7”
combination 18 21 23 24 26 (equivalent to cluster 15 without data set 25) points to directory “lysozyme/blend_8”
combination 18 21 23 24 25 (equivalent to cluster 15 without data set 26) points to directory “lysozyme/blend_9”
Final statistics can be viewed in the "MERGED_STATISTICS.info" file under "lysozyme/blend_9/combined_files" or simply using the interface:
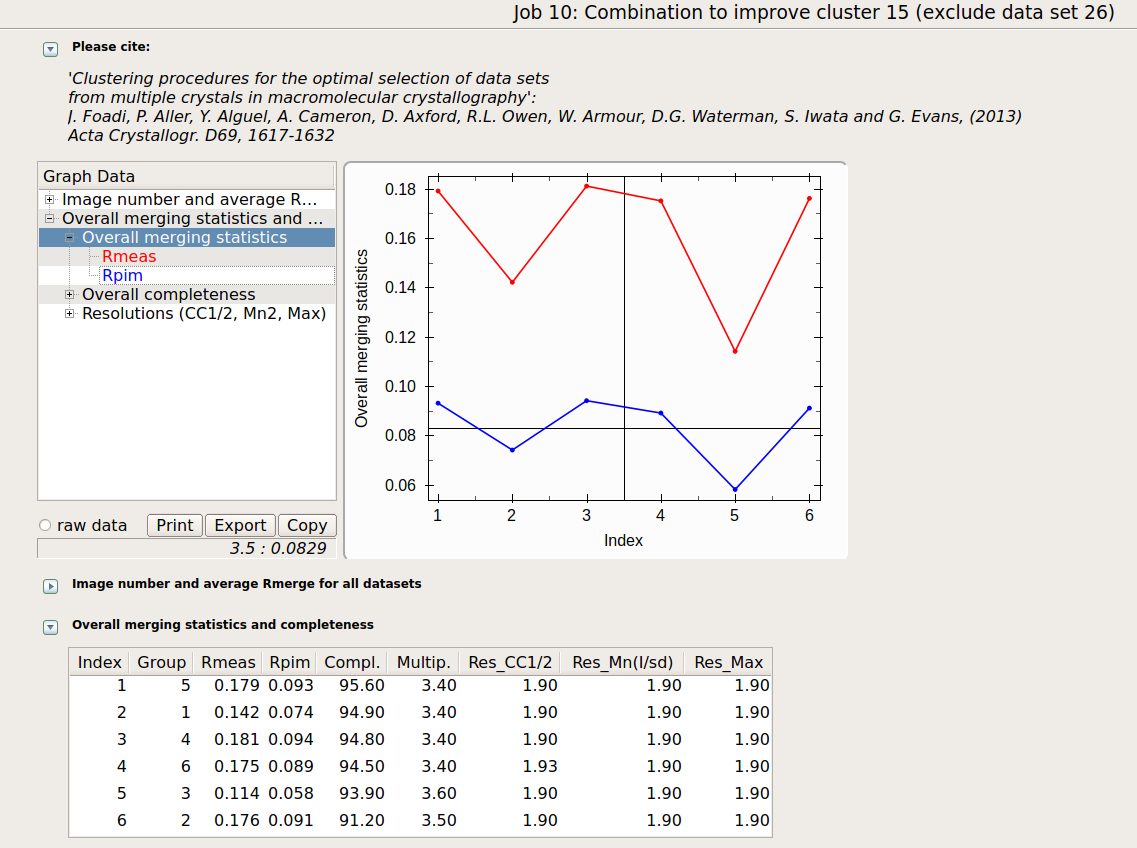
The lowest Rmeas is the one for group 3. Thus, discarding sweep 23 from cluster 15 causes Rmeas to improve from 0.161 to 0.114, while completeness decrease a little from 96.0% to 93.9%. This is essentially cluster 11; our initial guess pointing at data set 23 as the one making things worst turned out to be correct. Similar combinations can be tried on all other clusters. Results are shown in the following picture:
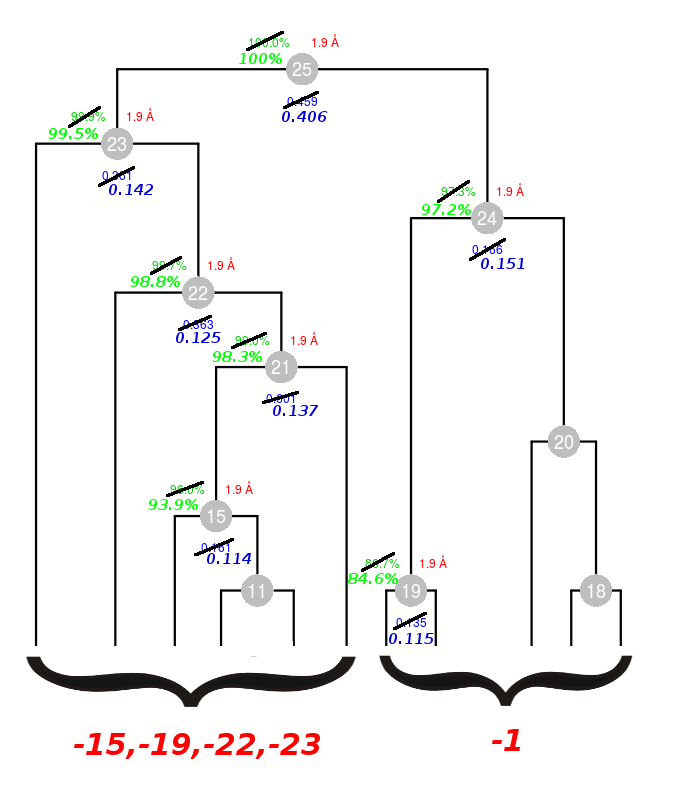
Crossed values have been updated with values calculated with the new combinations. For example cluster 15 had Rmeas=0.161 and this becomes 0.114 after filtering. At the same time its completeness drops from 96.0% to 93.9%. In total 5 data sets have been filtered out, 15, 19, 22, 23 from the left branch and 1 from the right branch. Statistics look much better for each branch. Their union into cluster 25 still yields a high value of Rmeas because the two groups of crystals, as we know, are non isomorphous in reason of the bromine. Try and reproduce the same results by executing BLEND in combination mode.
A final remark: all scaled mtz files do not contain structure factors, but only scaled intensities. To create structure factors and proceed with phasing, model building and refinement, you need to run the CCP4 program TRUNCATE or any other software that does an equivalent job.
Easy SAD phasing with multiple crystals
Each one of these data sets is not complete. The anomalous signal produced by the heavy bromine atoms can be detected
even with partial data sets, but effective phasing can only happen reliably with good anomalous completeness. For this
reason it is necessary to merge data from these multiple crystals to obtain full anomalous completeness. The resulting
high redundancy makes the anomalous signal even stronger, provided isomorphism is conserved.
In this tutorial we will use BLEND to first explore resolution and completeness of data and then to obtain complete
data sets that will be subsequently used to find the heavy atom substructure. The CCP4 GUI will not be used, so that the
whole tutorial is simplified, but the GUI should be used if you are unfamiliar with command line execution.
First let us create a new directory, called "lysBr" inside the "blend_tutorial" directory:
mkdir lysBr
Data in this case are stored in directory "lysBr" under "blend_tutorial/data". But they are spread over 12 different directories. In order to prepare data for BLEND we have two options; either copy all data into a single directory, or edit a file with paths for all 12 input files. In this case the second option is better because all individual input files have been produced by XDS and have identical name, "INTEGRATE.HKL". A quick way to prepare the input file for BLEND is to use the unix command "find". First let us move inside the "lysBr" directory (our working directory, in this case):
cd lysBr
Next, use "find" starting from "data/lysBr" directory and re-direct the results into a new file called "original.dat". This file is the initial input for BLEND (the "sort" command after the pipe symbol "|" is to sort directory names; it's not needed in general, as input file order really does not matter in BLEND, but here it helps the tutorial description):
find ../data/lysBr -name "INTEGRATE.HKL" | sort > original.dat
The contents of "original.dat" should look like the following:
../data/lysBr/xtl001/INTEGRATE.HKL
../data/lysBr/xtl002/INTEGRATE.HKL
../data/lysBr/xtl003/INTEGRATE.HKL
../data/lysBr/xtl004/INTEGRATE.HKL
../data/lysBr/xtl005/INTEGRATE.HKL
../data/lysBr/xtl006/INTEGRATE.HKL
../data/lysBr/xtl009/INTEGRATE.HKL
../data/lysBr/xtl011/INTEGRATE.HKL
../data/lysBr/xtl012/INTEGRATE.HKL
../data/lysBr/xtl015/INTEGRATE.HKL
../data/lysBr/xtl016/INTEGRATE.HKL
../data/lysBr/xtl017/INTEGRATE.HKL
BLEND can now be executed in analysis mode using "original.dat" as input (remember to hit the "enter" key twice, as we hav not prepared any keywords file):
blend -a original.dat
The tree produced should look like the following:
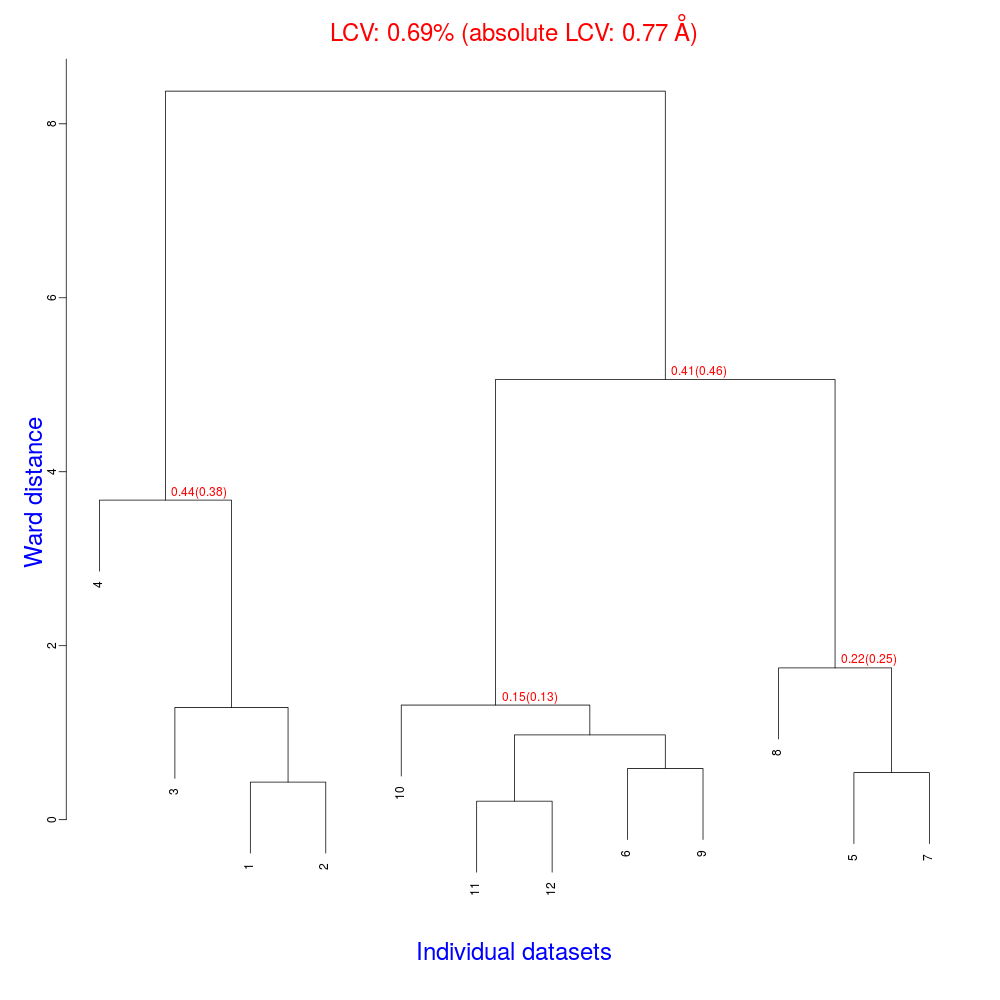
The linear cell variation for all crystals has a small 0.69% value, which means all crystals are fairly isomorphous. Among all files produced by BLEND in the present working directory the new "xds_files" directory and "xds_lookup_table.txt" file have been produced. This happens every time input data come from XDS integration files. BLEND makes use of POINTLESS to turn them into mtz files. All new files, including POINTLESS log files, are included in the "xds_files" directory. The "xds_lookup_table.txt" helps users to keep track of file origin, because all XDS files are re-named in the process. In this specific case, for example, directory "xds_files" includes the following files:
dataset_001.mtz
dataset_002.mtz
dataset_003.mtz
dataset_004.mtz
dataset_005.mtz
dataset_006.mtz
dataset_007.mtz
dataset_008.mtz
dataset_009.mtz
dataset_010.mtz
dataset_011.mtz
dataset_012.mtz
pointless_001.log
pointless_002.log
pointless_003.log
pointless_004.log
pointless_005.log
pointless_006.log
pointless_007.log
pointless_008.log
pointless_009.log
pointless_010.log
pointless_011.log
pointless_012.log
The associated "xds_lookup_table.txt" looks like the following:
dataset_001.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl001/INTEGRATE.HKL
dataset_002.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl002/INTEGRATE.HKL
dataset_003.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl003/INTEGRATE.HKL
dataset_004.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl004/INTEGRATE.HKL
dataset_005.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl005/INTEGRATE.HKL
dataset_006.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl006/INTEGRATE.HKL
dataset_007.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl009/INTEGRATE.HKL
dataset_008.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl011/INTEGRATE.HKL
dataset_009.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl012/INTEGRATE.HKL
dataset_010.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl015/INTEGRATE.HKL
dataset_011.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl016/INTEGRATE.HKL
dataset_012.mtz /home/james/safe-test/workBLEND/blend_tutorial/data/lysBr/xtl017/INTEGRATE.HKL
In order to estimate data resolutions we can run BLEND in combination mode on each individual data set. Strictly speaking this is not needed, and merged data corresponding to all clusters can be calculated straight away, but exploring individual data sets (in those instances where data wedges and symmetry allow scaling) can help to figure out data quality. We know that the space group for these samples of lysozyme is P 43 21 2 and can fix this in the keywords file. Thus, a new file, named "bkeys.dat" can be edited to include the following line:
CHOOSE SPACEGROUP P43212
Next, BLEND is executed in combination mode on all 12 data sets:
blend -c 1 < bkeys.dat
blend -c 2 < bkeys.dat
blend -c 3 < bkeys.dat
blend -c 4 < bkeys.dat
blend -c 5 < bkeys.dat
blend -c 6 < bkeys.dat
blend -c 7 < bkeys.dat
blend -c 8 < bkeys.dat
blend -c 9 < bkeys.dat
blend -c 10 < bkeys.dat
blend -c 11 < bkeys.dat
blend -c 12 < bkeys.dat
All files produced by these 12 runs of BLEND are contained in a directory called "combined_files". Let us rename this directory as "individual_files" because it includes process data from individual data sets:
mv combined_files individual_files
File "MERGING_STATISTICS.info" inside "individual_files" is a summary of overall statistics:
Group
Comple-
Multi-
Reso
Reso
Reso
Number
Rmeas
Rpim
teness
plicity
CC1/2
Mn(I/sd)
Max
1 0.066 0.043 58.00 1.50 1.47 1.47 1.47
2 0.071 0.040 55.50 2.50 1.40 1.40 1.40
3 0.294 0.112 92.60 5.40 1.64 1.75 1.43
4 0.083 0.047 67.80 2.20 1.50 1.50 1.50
5 0.067 0.044 71.40 1.50 1.39 1.39 1.39
6 0.206 0.129 84.40 2.00 1.64 1.83 1.53
7 0.191 0.116 70.50 2.10 1.50 1.50 1.50
8 0.050 0.032 52.40 1.80 1.48 1.48 1.48
9 0.127 0.087 72.30 1.50 1.49 1.49 1.49
10 0.556 0.385 67.10 1.40 1.92 2.03 1.49
11 0.082 0.054 60.00 1.70 1.40 1.40 1.40
12 0.074 0.045 71.30 2.10 1.39 1.39 1.39
From the "Completeness" column we can see that the only complete data set is data set 3. Its resolution, as suggested by the "Reso CC1/2" and "Reso Mn(I/sd)" columns falls between 1.64 Å and 1.75 Å. Resolutions are higher for the other partial data sets and it seems reasonable to adopt 1.6 Å as a common resolution for all data. The previous table can be reproduced using this resolution if the keywords file includes the following lines ("ANOMALOUS ON" has been added to make sure AIMLESS treats Friedel's opposites separately and that it returns the corresponding anomalous statistics):
CHOOSE SPACEGROUP P43212
RESO HIGH 1.6
ANOMALOUS ON
The corresponding table is:
Group
Comple-
Multi-
Reso
Reso
Reso
Number
Rmeas
Rpim
teness
plicity
CC1/2
Mn(I/sd)
Max
1 0.063 0.041 59.40 1.60 1.60 1.60 1.60
2 0.068 0.039 60.60 2.60 1.60 1.60 1.60
3 0.213 0.078 94.70 5.80 1.63 1.76 1.60
4 0.077 0.043 67.90 2.20 1.60 1.60 1.60
5 0.063 0.042 79.60 1.60 1.60 1.60 1.60
6 0.191 0.120 84.80 2.00 1.64 1.83 1.60
7 0.175 0.106 70.90 2.20 1.60 1.60 1.60
8 0.048 0.030 52.70 1.90 1.60 1.60 1.60
9 0.123 0.085 75.40 1.50 1.60 1.60 1.60
10 0.467 0.320 70.00 1.50 1.88 2.04 1.60
11 0.079 0.052 68.70 1.80 1.60 1.60 1.60
12 0.071 0.044 79.30 2.10 1.60 1.60 1.60
The newly created "combined_files" directory can be renamed "individual_files_1.6", to differentiate it from "individual_files":
mv combined_files individual_files_1.6
Next, BLEND is executed in synthesis mode using the same keywords file, to produce merged data for all nodes of the dendrogram at this resolution. As a value of 10 is high enough to include all nodes of the dendrogram, the resulting command line will result as follows:
blend -s 10 < bkeys.dat > blend_synthesis.log
All results are, this time, dumped in directory "merged_files". We can appreciate overall statistics by viewing the "MERGING_STATISTICS.info" file inside this directory. These statistics, though, give no information we can use to judge the quality of data for phasing. This information is still not extracted and organised by BLEND because the whole issue is currently under investigation. Future addition in "MERGING_STATISTICS.info" will include such information. For now we can extract specific lines from all AIMLESS logs using the unix "grep" command. For instance, anomalous completeness is important to judge the quality of anomalous data. We can extract it with the following unix line:
grep "Anomalous completeness " merged_files/aimless*.log
The output produced is the following:
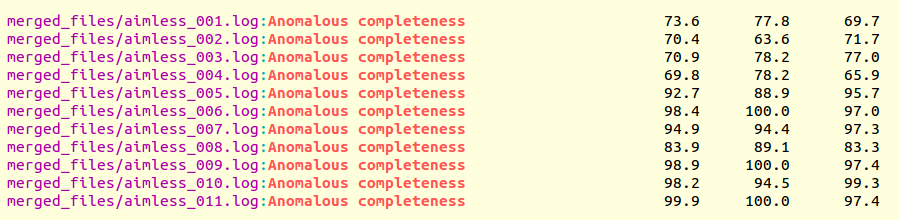
Clusters with satisfactory anomalous completeness are 5, 6, 7, 9, 10, 11 (greater than 90%). These are data sets on which we can try the procedure to find heavy atoms substructure. These files can be compared with the only individual data set with the highest anomalous completeness, data set 3. A "grep" line similar to the one previously used can show this:
grep "Anomalous completeness " individual_files_1.6/aimless*.log
Resulting output:
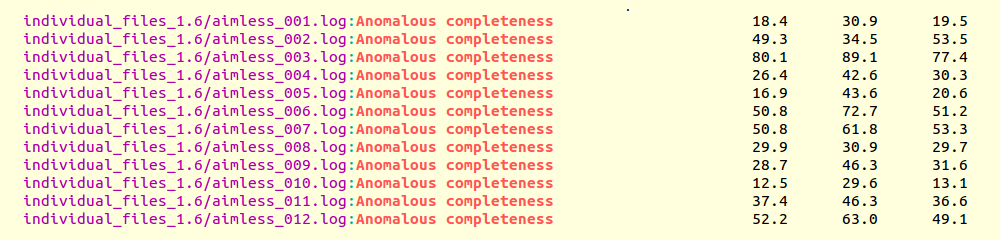
The value 80.1% for the anomalous completeness of data set 3 is not close to 90%, but the only decent one
obtainable as indvidual data set, and it is with this one that all merged ones will have to be compared. The message
to be taken home is that when only partial data are available it is still possible to combine them and obtain a complete data set,
rather than wait for new crystals and another data collection. Furthermore, the redundancy implied by merging data together can
increase the anomalous signal and, thus, strengthen the phasing power.
In what follows we have used the SHELX family of programs to prepare data, measure anomalous signal and find the heavy
atoms substructure (Sheldrick (2010)). These programs work with input files of a specific format ("sca" files).
In order to prepare these files we need, first, to turn scaled intensities into structure factors using the CCP4 program TRUNCATE.
Then we use "MTZ2SCA" to turn mtz files into sca files. Everything will be executed inside a new directory called
"anoproc". In this directory we need to create seven more directories, one for each data set to be processed. The whole series of
unix commands to be typed is:
mkdir anoproc
cd anoproc
mkdir lysBr_ind_003
mkdir lysBr_ind_012
mkdir lysBr_cl_005
mkdir lysBr_cl_006
mkdir lysBr_cl_007
mkdir lysBr_cl_009
mkdir lysBr_cl_010
mkdir lysBr_cl_011
The procedure to produce sca files ready to be used in SHELX will be now shown only for one of the data sets involved, the one corresponding to cluster 6, but it is meant to be repeated in exactly the same way for all other data sets. First we move to directory "lysBr_cl_006". Then we run TRUNCATE, using command lines, in the following way:
cd lysBr_cl_006
truncate HKLIN ../../merged_files/scaled_006.mtz HKLOUT truncate_cl_006.mtz
After having entered this line execution halts as TRUNCATE expects some keywords. We can use default values and just type "END" (and hit the "Enter" key). After execution a file "truncate_cl_006.mtz" should appear in the directory. To translate this file into a sca file we use MTZ2SCA in a very simple way:
mtz2sca truncate_cl_006.mtz
Now also "truncate_cl_006.sca" should be present in the directory. As mentioned before, the procedure just applied on data set 6 should be repeated for the other 6 data sets in the "anoproc" directory. Once all sca files have been obtained, programs SHELXC and SHELXD can be executed to obtain the heavy atoms substructure. The heavy atom for this specific case is bromine (BR - 8 sites) and we have tried 1000 trials in SHELXD. Results are tabulated as follows:
| Data set | Sufficient d"/sig ? | Solution ? | CCall | CCweak | Cfom |
| ind_003 | yes (-) | yes (-) | 31.04 | 16.85 | 47.89 |
| cl_005 | yes | yes | 42.82 | 26.07 | 68.89 |
| cl_006 | yes | yes | 36.24 | 20.73 | 56.96 |
| cl_007 | yes | yes | 44.07 | 27.38 | 71.44 |
| cl_009 | yes | yes | 32.74 | 19.55 | 52.28 |
| cl_010 | yes | yes | 43.03 | 26.80 | 69.83 |
| cl_011 | yes | yes | 41.56 | 26.83 | 68.38 |
The phasing power of bromine in a stable structure like lysozyme is clear from the relatively high CCall (> 30) for the individual data set "ind_003". It is also clear, though, that the anomalous signal becomes much more effective when multiple isomorphous crystals are merged together. Complete clusters which includes "ind_003", i.e. cluster 006 and 009, improve the anomalous signal of "ind_003" alone. Clusters 005, 007 and 010 perform better than the other clusters.
Automated filtering of lysozyme crystals using the "cF" variant of the combination mode
Filtering for the 28 sweeps of lysozyme crystals was carried out manually in Tutorial 2. There BLEND was executed in combination mode and filtering was done trying many combinations and selecting the ones giving improved Rmeas values. It is also possible to let the program carry out filtering in automated way, using the "F" variant of combination mode (option "-cF"). A starting cluster is first indicated by the user. This cluster will have to be redundant because filtering means "throwing away" data sets, and this implies potentially decreasing completeness. The user will have to add to the keywords file the final completeness that can be tolerated (default 95%), as well as the maximum number of iteration cycles (default 5). The program will, then, discard, at every cycle, the single data set having largest overall Rmerge.
Let's create a new directory for this tutorial, say "lysozyme2", and execute BLEND in analysis mode, pointing to directory "data/lysozyme". Let's also create a keyword file, "bkeys.dat", with the following lines:
RESO HIGH 1.9
TOLERANCE 100
COMPLETENESS 90
MAXCYCLE 20
where the resolution has been fixed at 1.9 Å to enable comparison with the filtering done at Tutorial 2. Here we are also asking for completeness not to go below 90% and have increased the number of possible cycles to 20, so to make it possible to filter out several data sets. Thus, for the analysis mode:
blend -a ../data/lysozyme < bkeys.dat > blend_analysis.log
If we look at the dendrogram we notice that data sets 19 and 21 are very different from the other data sets. Thus, like done in Tutorial 2, we copy file "mtz_names.dat" into a new file named "original.dat" and delete lines 19 and 21. Next we re-run BLEND in analysis mode as follows:
blend -a original.dat < bkeys.dat > blend_analysis.log
The isomorphism, as measured by aLCV, for all clusters in this group of 28 data sets, can be displayed with a plot produced using BLEND in graphics mode, with the type "DO":
blend -g DO 27 10
The plot produced can be found in sub-directory "graphics", and it is displayed here:
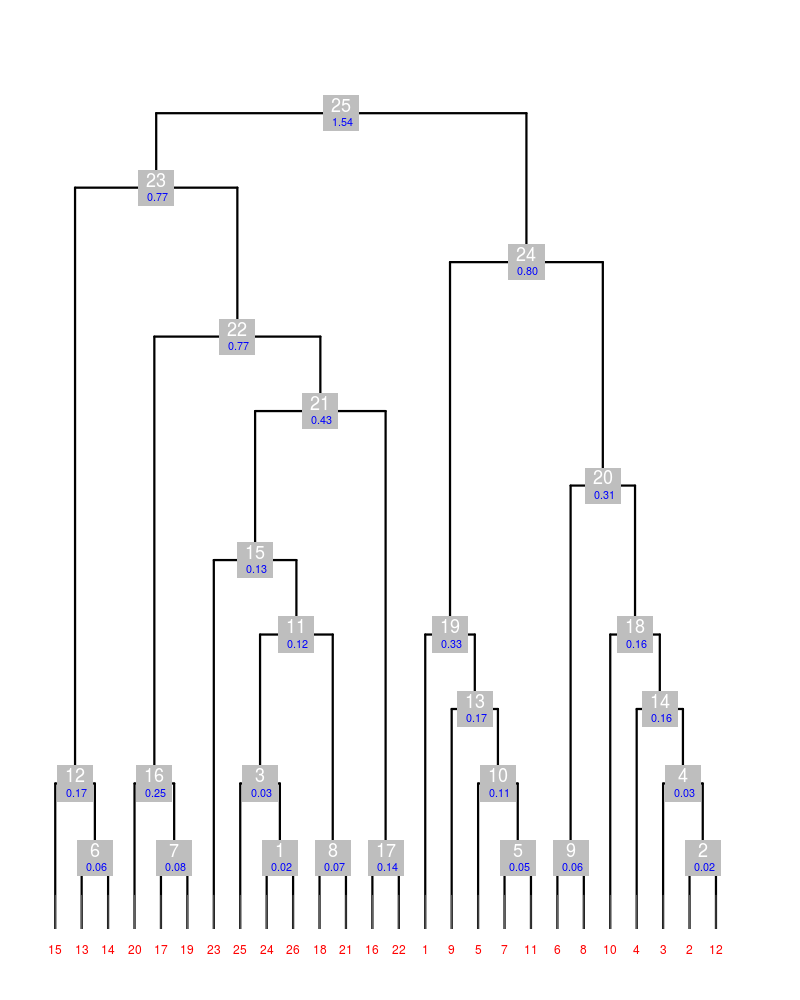
Both left (cluster 23) and right (cluster 24) branches of the dendrogram have relatively small aLCV values. This means that all crystals in each branch can be considered isomorphous. We can, then, delete some of the data sets in each branch so to bring Rmeas (or, even better, Rpim) down. As mentioned above, this is done using the option "-cF". In order to compare results with the cluster before the filtering we will also run BLEND in simple combination mode. Thus, for the left branch, we use:
blend -c [23] < bkeys.dat > blend_synthesis_23.log
blend -cF [23] < bkeys.dat > blend_synthesisF_23.log
Results for both runs can be found in the directory "combined_files". More specifically, file "MERGING_STATISTICS.info" appears as follows:
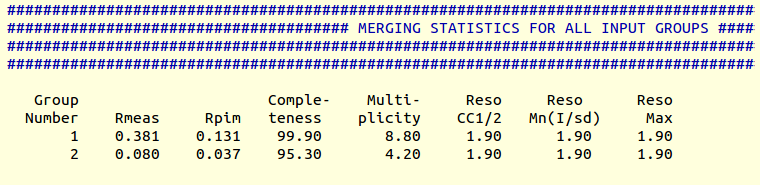
By removing several data sets, Rpim has dropped from 0.131 to 0.037. Completeness has dropped to 95.3%. Only 10 cycles were needed for taking the statistics to this level; cycle 11 would have taken completeness to 82.9%, lower than our target 90%. This is why cycling was interrupted by the program. More information can be found in the log file. We can repeat the same exercise for the right branch (cluster 24):
blend -c [24] < bkeys.dat > blend_synthesis_24.log
blend -cF [24] < bkeys.dat > blend_synthesisF_24.log
This time results are given, in file "MERGING_STATISTICS.info", by the third and fourth group:
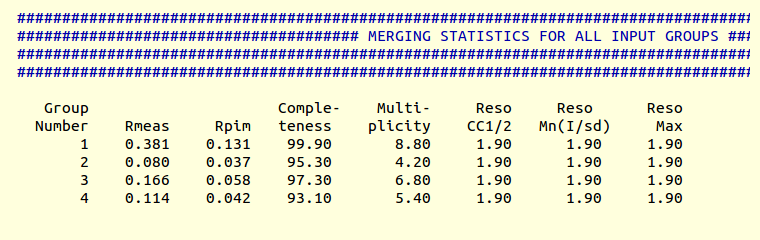
This time the Rpim dropped from 0.166 to 0.114, completeness went to 93.1% and the job terminated after 3 cycles as the fourth cycle would have taken completeness down to 89%, below the 90% set threshold.
We could have started from the top cluster and removed automatically data sets down to the desired completeness. But in so doing it is also possible to remove information which is potentially useful. Thus, this variant of the combination mode must be used cautiuosly.
(2010) G. M. Sheldrick, "Experimental phasing with SHELXC/D/E: combining chain tracing with density modification", Acta Cryst. D66, 479-485
(2011) Q. Liu, Z. Zhang and W.A. Hendrickson, "Multi-crystal anomalous diffraction for low-resolution macromolecular phasing", Acta Cryst. D67, 45-49
(2012) R. Giordano, R.M.F. Leal, G.P. Bourenkov, S. McSweeney and A.N. Popov, “The application of hierarchical cluster analysis to the selection of isomorphous crystals", Acta Cryst. D68, 649-658
(2012) D. Axford, R.L. Owen, J. Aishima, J. Foadi, A.W. Morgan, J.I. Robinson, J.E. Nettleship, R.J. Owens, I. Moraes, E.E. Fry, J.M. Grimes, K. Harlos, A. Kotecha, J. Ren, G. Sutton, T.S. Walter, D.I. Stuart and G. Evans, "In situ macromolecular crystallography using microbeams", Acta Cryst. D68, 592-600
(2012) M.A. Hanson, C.B. Roth, E. Jo, M.T. Griffith, F.L. Scott, G. Reinhart, H. Desale, B. Clemons, S.M. Cahalan, S.C. Schuerer, M.G. Sanna, G.W. Han, P. Kuhn, H. Rosen, R.C. Stevens, "Crystal structure of a lipid G protein-coupled receptor"
(2013) J. Foadi, P. Aller, Y. Alguel, A. Cameron, D. Axford, R.L. Owen, W. Armour, D.G. Waterman, S. Iwata and G. Evans, "Clustering procedures for the optimal selection of data sets from multiple crystals in macromolecular crystallography", Acta Cryst. D69, 1617-1632, Science 335, 851-855